Genome maintenance in the context of 4D chromatin condensation
- PMID: 27098512
- PMCID: PMC4956502
- DOI: 10.1007/s00018-016-2221-2
Genome maintenance in the context of 4D chromatin condensation
Abstract
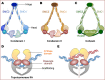
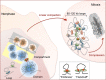
The eukaryotic genome is packaged in the three-dimensional nuclear space by forming loops, domains, and compartments in a hierarchical manner. However, when duplicated genomes prepare for segregation, mitotic cells eliminate topologically associating domains and abandon the compartmentalized structure. Alongside chromatin architecture reorganization during the transition from interphase to mitosis, cells halt most DNA-templated processes such as transcription and repair. The intrinsically condensed chromatin serves as a sophisticated signaling module subjected to selective relaxation for programmed genomic activities. To understand the elaborate genome-epigenome interplay during cell cycle progression, the steady three-dimensional genome requires a time scale to form a dynamic four-dimensional and a more comprehensive portrait. In this review, we will dissect the functions of critical chromatin architectural components in constructing and maintaining an orderly packaged chromatin environment. We will also highlight the importance of the spatially and temporally conscious orchestration of chromatin remodeling to ensure high-fidelity genetic transmission.
Keywords: Cancer; Cell cycle; Chromatin architecture; Epigenome; Genome stability.
Figures
Similar articles
-
An insight into understanding the coupling between homologous recombination mediated DNA repair and chromatin remodeling mechanisms in plant genome: an update.Cell Cycle. 2021 Sep;20(18):1760-1784. doi: 10.1080/15384101.2021.1966584. Epub 2021 Aug 26. Cell Cycle. 2021. PMID: 34437813 Free PMC article. Review.
-
Role of histone phosphorylation in chromatin dynamics and its implications in diseases.Subcell Biochem. 2007;41:319-36. Subcell Biochem. 2007. PMID: 17484134 Review.
-
Epigenetics and genome stability.Mamm Genome. 2020 Jun;31(5-6):181-195. doi: 10.1007/s00335-020-09836-2. Epub 2020 Apr 15. Mamm Genome. 2020. PMID: 32296924 Review.
-
Chromatin replication and epigenome maintenance.Nat Rev Mol Cell Biol. 2012 Feb 23;13(3):153-67. doi: 10.1038/nrm3288. Nat Rev Mol Cell Biol. 2012. PMID: 22358331 Review.
-
Emerging roles of linker histones in regulating chromatin structure and function.Nat Rev Mol Cell Biol. 2018 Mar;19(3):192-206. doi: 10.1038/nrm.2017.94. Epub 2017 Oct 11. Nat Rev Mol Cell Biol. 2018. PMID: 29018282 Free PMC article. Review.
Cited by
-
Episomal HBV persistence within transcribed host nuclear chromatin compartments involves HBx.Epigenetics Chromatin. 2018 Jun 22;11(1):34. doi: 10.1186/s13072-018-0204-2. Epigenetics Chromatin. 2018. PMID: 29933745 Free PMC article.
-
Circulating Nucleosomes and Nucleosome Modifications as Biomarkers in Cancer.Cancers (Basel). 2017 Jan 8;9(1):5. doi: 10.3390/cancers9010005. Cancers (Basel). 2017. PMID: 28075351 Free PMC article. Review.
-
Differentially accessible, single copy sequences form contiguous domains along metaphase chromosomes that are conserved among multiple tissues.Mol Cytogenet. 2021 Oct 20;14(1):49. doi: 10.1186/s13039-021-00567-w. Mol Cytogenet. 2021. PMID: 34670606 Free PMC article.
-
A Paradigm Revolution or Just Better Resolution-Will Newly Emerging Superresolution Techniques Identify Chromatin Architecture as a Key Factor in Radiation-Induced DNA Damage and Repair Regulation?Cancers (Basel). 2020 Dec 23;13(1):18. doi: 10.3390/cancers13010018. Cancers (Basel). 2020. PMID: 33374540 Free PMC article. Review.
-
Cohesin-Dependent and -Independent Mechanisms Mediate Chromosomal Contacts between Promoters and Enhancers.Cell Rep. 2020 Jul 21;32(3):107929. doi: 10.1016/j.celrep.2020.107929. Cell Rep. 2020. PMID: 32698000 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

