Machine-learning scoring functions to improve structure-based binding affinity prediction and virtual screening
- PMID: 27110292
- PMCID: PMC4832270
- DOI: 10.1002/wcms.1225
Machine-learning scoring functions to improve structure-based binding affinity prediction and virtual screening
Abstract
Docking tools to predict whether and how a small molecule binds to a target can be applied if a structural model of such target is available. The reliability of docking depends, however, on the accuracy of the adopted scoring function (SF). Despite intense research over the years, improving the accuracy of SFs for structure-based binding affinity prediction or virtual screening has proven to be a challenging task for any class of method. New SFs based on modern machine-learning regression models, which do not impose a predetermined functional form and thus are able to exploit effectively much larger amounts of experimental data, have recently been introduced. These machine-learning SFs have been shown to outperform a wide range of classical SFs at both binding affinity prediction and virtual screening. The emerging picture from these studies is that the classical approach of using linear regression with a small number of expert-selected structural features can be strongly improved by a machine-learning approach based on nonlinear regression allied with comprehensive data-driven feature selection. Furthermore, the performance of classical SFs does not grow with larger training datasets and hence this performance gap is expected to widen as more training data becomes available in the future. Other topics covered in this review include predicting the reliability of a SF on a particular target class, generating synthetic data to improve predictive performance and modeling guidelines for SF development. WIREs Comput Mol Sci 2015, 5:405-424. doi: 10.1002/wcms.1225 For further resources related to this article, please visit the WIREs website.
Figures

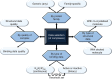
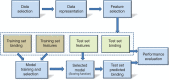

References
-
- Schneider G. Virtual screening: an endless staircase? Nat Rev Drug Discov 2010, 9:273–276. - PubMed
-
- Vasudevan SR, Churchill GC. Mining free compound databases to identify candidates selected by virtual screening. Expert Opin Drug Discov 2009, 4:901–906. - PubMed
-
- Villoutreix BO, Renault N, Lagorce D, Sperandio O, Montes M, Miteva MA. Free resources to assist structure‐based virtual ligand screening experiments. Curr Protein Pept Sci 2007, 8:381–411. - PubMed
-
- Xing L, McDonald JJ, Kolodziej SA, Kurumbail RG, Williams JM, Warren CJ, O'Neal JM, Skepner JE, Roberds SL. Discovery of potent inhibitors of soluble epoxide hydrolase by combinatorial library design and structure‐based virtual screening. J Med Chem 2011, 54:1211–1222. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
