RpiRc Is a Pleiotropic Effector of Virulence Determinant Synthesis and Attenuates Pathogenicity in Staphylococcus aureus
- PMID: 27113358
- PMCID: PMC4936357
- DOI: 10.1128/IAI.00285-16
RpiRc Is a Pleiotropic Effector of Virulence Determinant Synthesis and Attenuates Pathogenicity in Staphylococcus aureus
Abstract
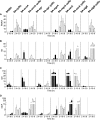
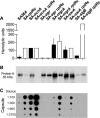
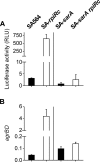
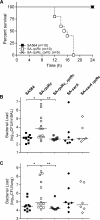
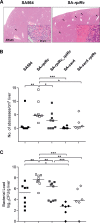
In Staphylococcus aureus, metabolism is intimately linked with virulence determinant biosynthesis, and several metabolite-responsive regulators have been reported to mediate this linkage. S. aureus possesses at least three members of the RpiR family of transcriptional regulators. Of the three RpiR homologs, RpiRc is a potential regulator of the pentose phosphate pathway, which also regulates RNAIII levels. RNAIII is the regulatory RNA of the agr quorum-sensing system that controls virulence determinant synthesis. The effect of RpiRc on RNAIII likely involves other regulators, as the regulators that bind the RNAIII promoter have been intensely studied. To determine which regulators might bridge the gap between RpiRc and RNAIII, sarA, sigB, mgrA, and acnA mutations were introduced into an rpiRc mutant background, and the effects on RNAIII were determined. Additionally, phenotypic and genotypic differences were examined in the single and double mutant strains, and the virulence of select strains was examined using two different murine infection models. The data suggest that RpiRc affects RNAIII transcription and the synthesis of virulence determinants in concert with σ(B), SarA, and the bacterial metabolic status to negatively affect virulence.
Copyright © 2016, American Society for Microbiology. All Rights Reserved.
Figures

Similar articles
-
Staphylococcus aureus Coordinates Leukocidin Expression and Pathogenesis by Sensing Metabolic Fluxes via RpiRc.mBio. 2016 Jun 21;7(3):e00818-16. doi: 10.1128/mBio.00818-16. mBio. 2016. PMID: 27329753 Free PMC article.
-
RpiR homologues may link Staphylococcus aureus RNAIII synthesis and pentose phosphate pathway regulation.J Bacteriol. 2011 Nov;193(22):6187-96. doi: 10.1128/JB.05930-11. Epub 2011 Sep 16. J Bacteriol. 2011. PMID: 21926234 Free PMC article.
-
Rat/MgrA, a regulator of autolysis, is a regulator of virulence genes in Staphylococcus aureus.Infect Immun. 2005 Mar;73(3):1423-31. doi: 10.1128/IAI.73.3.1423-1431.2005. Infect Immun. 2005. PMID: 15731040 Free PMC article.
-
Staphylococcus aureus RNAIII and Its Regulon Link Quorum Sensing, Stress Responses, Metabolic Adaptation, and Regulation of Virulence Gene Expression.Annu Rev Microbiol. 2016 Sep 8;70:299-316. doi: 10.1146/annurev-micro-102215-095708. Epub 2016 Jul 6. Annu Rev Microbiol. 2016. PMID: 27482744 Review.
-
Regulation of Staphylococcus aureus Virulence.Microbiol Spectr. 2019 Apr 5;7(2):10.1128/microbiolspec.gpp3-0031-2018. doi: 10.1128/microbiolspec.GPP3-0031-2018. Microbiol Spectr. 2019. PMID: 30953424 Free PMC article. Review.
Cited by
-
PtpA, a secreted tyrosine phosphatase from Staphylococcus aureus, contributes to virulence and interacts with coronin-1A during infection.J Biol Chem. 2018 Oct 5;293(40):15569-15580. doi: 10.1074/jbc.RA118.003555. Epub 2018 Aug 21. J Biol Chem. 2018. PMID: 30131335 Free PMC article.
-
Proteomic Adaptation of Clostridioides difficile to Treatment with the Antimicrobial Peptide Nisin.Cells. 2021 Feb 11;10(2):372. doi: 10.3390/cells10020372. Cells. 2021. PMID: 33670309 Free PMC article.
-
RNAIII is linked with the pentose phosphate pathway through the activation of RpiRc in Staphylococcus aureus.mSphere. 2024 May 29;9(5):e0034823. doi: 10.1128/msphere.00348-23. Epub 2024 Apr 9. mSphere. 2024. PMID: 38591898 Free PMC article.
-
Differential evolution in 3'UTRs leads to specific gene expression in Staphylococcus.Nucleic Acids Res. 2020 Mar 18;48(5):2544-2563. doi: 10.1093/nar/gkaa047. Nucleic Acids Res. 2020. PMID: 32016395 Free PMC article.
-
The Phosphoarginine Phosphatase PtpB from Staphylococcus aureus Is Involved in Bacterial Stress Adaptation during Infection.Cells. 2021 Mar 14;10(3):645. doi: 10.3390/cells10030645. Cells. 2021. PMID: 33799337 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

