A Multi-Site Study of Norovirus Molecular Epidemiology in Australia and New Zealand, 2013-2014
- PMID: 27116221
- PMCID: PMC4846056
- DOI: 10.1371/journal.pone.0145254
A Multi-Site Study of Norovirus Molecular Epidemiology in Australia and New Zealand, 2013-2014
Abstract
Background: Norovirus (NoV) is the major cause of acute gastroenteritis across all age groups. In particular, variants of genogroup II, genotype 4 (GII.4) have been associated with epidemics globally, occurring approximately every three years. The pandemic GII.4 variant, Sydney 2012, was first reported in early 2012 and soon became the predominant circulating NoV strain globally. Despite its broad impact, both clinically and economically, our understanding of the fundamental diversity and mechanisms by which new NoV strains emerge remains limited. In this study, we describe the molecular epidemiological trends of NoV-associated acute gastroenteritis in Australia and New Zealand between January 2013 and June 2014.
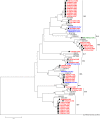
Methodology: Overall, 647 NoV-positive clinical faecal samples from 409 outbreaks and 238 unlinked cases of acute gastroenteritis were examined by RT-PCR and sequencing. Phylogenetic analysis was then performed to identify NoV capsid genotypes and to establish the temporal dominance of circulating pandemic GII.4 variants. Recombinant viruses were also identified based on analysis of the ORF1/2 overlapping region.
Findings: Peaks in NoV activity were observed, however the timing of these epidemics varied between different regions. Overall, GII.4 NoVs were the dominant cause of both outbreaks and cases of NoV-associated acute gastroenteritis (63.1%, n = 408/647), with Sydney 2012 being the most common GII.4 variant identified (98.8%, n = 403/408). Of the 409 reported NoV outbreaks, aged-care facilities were the most common setting in both Western Australia (87%, n = 20/23) and New Zealand (58.1%, n = 200/344) while most of the NoV outbreaks were reported from hospitals (38%, n = 16/42) in New South Wales, Australia. An analysis of a subset of non-GII.4 viruses from all locations (125/239) showed the majority (56.8%, n = 71/125) were inter-genotype recombinants. These recombinants were surprisingly diverse and could be classified into 18 distinct recombinant types, with GII.P16/GII.13 (24% of recombinants) the most common.
Conclusion: This study revealed that following its emergence in 2012, GII.4 Sydney 2012 variant continued to be the predominant cause of NoV-associated acute gastroenteritis in Australia and New Zealand between 2013 and 2014.
Conflict of interest statement
Figures



References
-
- Estes MK, Prasad BV, Atmar RL. Noroviruses everywhere: has something changed? Curr Opin Infect Dis. 2006;19(5):467–74. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

