Seamless site-directed mutagenesis of the Saccharomyces cerevisiae genome using CRISPR-Cas9
- PMID: 27134651
- PMCID: PMC4850645
- DOI: 10.1186/s13036-016-0028-1
Seamless site-directed mutagenesis of the Saccharomyces cerevisiae genome using CRISPR-Cas9
Abstract
CRISPR assisted homology directed repair enables the introduction of virtually any modification to the Saccharomyces cerevisiae genome. Of obvious interest is the marker-free and seamless introduction of point mutations. To fulfill this promise, a strategy that effects single nucleotide changes while preventing repeated recognition and cutting by the gRNA/Cas9 complex is needed. We demonstrate a two-step method to introduce point mutations at 17 positions in the S. cerevisiae genome. We show the general applicability of the method, enabling the seamless introduction of single nucleotide changes at any location, including essential genes and non-coding regions. We also show a quantifiable phenotype for a point mutation introduced in gene GSH1. The ease and wide applicability of this general method, combined with the demonstration of its feasibility will enable genome editing at an unprecedented level of detail in yeast and other organisms.
Keywords: CRISPR-Cas9; Genome editing; Saccharomyces cerevisiae; Site-directed mutagenesis.
Figures
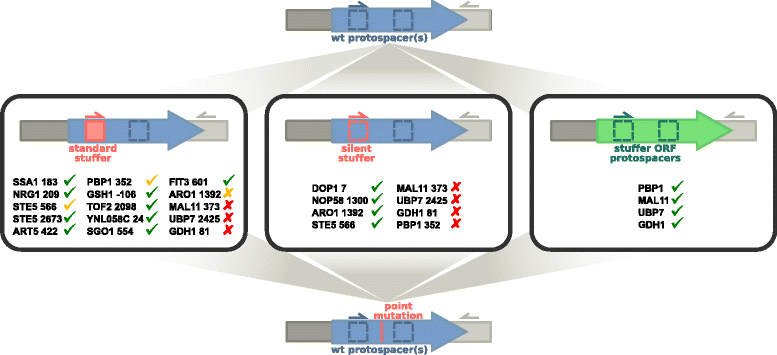
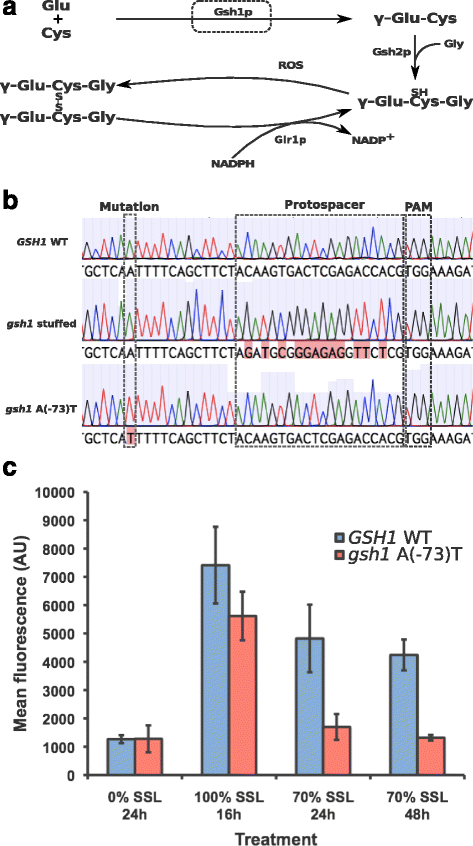
References
-
- Mans R, van Rossum HM, Wijsman M, Backx A, Kuijpers NGA, van den Broek M, Daran-Lapujade P, Pronk JT, van Maris AJA, Daran J-MG: CRISPR/Cas9: a molecular Swiss army knife for simultaneous introduction of multiple genetic modifications in Saccharomyces cerevisiae. FEMS Yeast Res. 2015; doi: 10.1093/femsyr/fov004 - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

