Development of Persister-FACSeq: a method to massively parallelize quantification of persister physiology and its heterogeneity
- PMID: 27142337
- PMCID: PMC4855238
- DOI: 10.1038/srep25100
Development of Persister-FACSeq: a method to massively parallelize quantification of persister physiology and its heterogeneity
Abstract
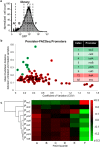
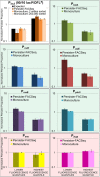
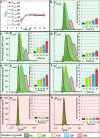
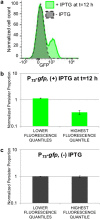
Bacterial persisters are thought to underlie the relapse of chronic infections. Knowledge of persister physiology would illuminate avenues for therapeutic intervention; however, such knowledge has remained elusive because persisters have yet to be segregated from other cell types to sufficient purity. This technical hurdle has stymied progress toward understanding persistence. Here we developed Persister-FACSeq, which is a method that uses fluorescence-activated cell sorting, antibiotic tolerance assays, and next generation sequencing to interrogate persister physiology and its heterogeneity. As a proof-of-concept, we used Persister-FACSeq on a library of reporters to study gene expression distributions in non-growing Escherichia coli, and found that persistence to ofloxacin is inversely correlated with the capacity of non-growing cells to synthesize protein. Since Persister-FACSeq can be applied to study persistence to any antibiotic in any environment for any bacteria that can harbor a fluorescent reporter, we anticipate that it will yield unprecedented knowledge of this detrimental phenotype.
Figures


Similar articles
-
Analyzing Persister Physiology with Fluorescence-Activated Cell Sorting.Methods Mol Biol. 2016;1333:83-100. doi: 10.1007/978-1-4939-2854-5_8. Methods Mol Biol. 2016. PMID: 26468102 Free PMC article.
-
Stationary-Phase Persisters to Ofloxacin Sustain DNA Damage and Require Repair Systems Only during Recovery.mBio. 2015 Sep 1;6(5):e00731-15. doi: 10.1128/mBio.00731-15. mBio. 2015. PMID: 26330511 Free PMC article.
-
Persisters: a distinct physiological state of E. coli.BMC Microbiol. 2006 Jun 12;6:53. doi: 10.1186/1471-2180-6-53. BMC Microbiol. 2006. PMID: 16768798 Free PMC article.
-
Formation, physiology, ecology, evolution and clinical importance of bacterial persisters.FEMS Microbiol Rev. 2017 May 1;41(3):219-251. doi: 10.1093/femsre/fux001. FEMS Microbiol Rev. 2017. PMID: 28333307 Review.
-
New-found fundamentals of bacterial persistence.Trends Microbiol. 2012 Dec;20(12):577-85. doi: 10.1016/j.tim.2012.08.009. Epub 2012 Sep 5. Trends Microbiol. 2012. PMID: 22959615 Review.
Cited by
-
Antibiotic Persistence as a Metabolic Adaptation: Stress, Metabolism, the Host, and New Directions.Pharmaceuticals (Basel). 2018 Feb 1;11(1):14. doi: 10.3390/ph11010014. Pharmaceuticals (Basel). 2018. PMID: 29389876 Free PMC article. Review.
-
Timing of DNA damage responses impacts persistence to fluoroquinolones.Proc Natl Acad Sci U S A. 2018 Jul 3;115(27):E6301-E6309. doi: 10.1073/pnas.1804218115. Epub 2018 Jun 18. Proc Natl Acad Sci U S A. 2018. PMID: 29915065 Free PMC article.
-
A High-Throughput Method for Identifying Novel Genes That Influence Metabolic Pathways Reveals New Iron and Heme Regulation in Pseudomonas aeruginosa.mSystems. 2021 Feb 2;6(1):e00933-20. doi: 10.1128/mSystems.00933-20. mSystems. 2021. PMID: 33531406 Free PMC article.
-
Microbial Metabolism Modulates Antibiotic Susceptibility within the Murine Gut Microbiome.Cell Metab. 2019 Oct 1;30(4):800-823.e7. doi: 10.1016/j.cmet.2019.08.020. Epub 2019 Sep 12. Cell Metab. 2019. PMID: 31523007 Free PMC article.
-
Recent Advances in Bacterial Persistence Mechanisms.Int J Mol Sci. 2023 Sep 20;24(18):14311. doi: 10.3390/ijms241814311. Int J Mol Sci. 2023. PMID: 37762613 Free PMC article. Review.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

