Gluten-specific antibodies of celiac disease gut plasma cells recognize long proteolytic fragments that typically harbor T-cell epitopes
- PMID: 27146306
- PMCID: PMC4857092
- DOI: 10.1038/srep25565
Gluten-specific antibodies of celiac disease gut plasma cells recognize long proteolytic fragments that typically harbor T-cell epitopes
Abstract
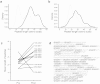
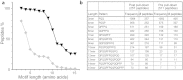
This study aimed to identify proteolytic fragments of gluten proteins recognized by recombinant IgG1 monoclonal antibodies generated from single IgA plasma cells of celiac disease lesions. Peptides bound by monoclonal antibodies in complex gut-enzyme digests of gluten treated with the deamidating enzyme transglutaminase 2, were identified by mass spectrometry after antibody pull-down with protein G beads. The antibody bound peptides were long deamidated peptide fragments that contained the substrate recognition sequence of transglutaminase 2. Characteristically, the fragments contained epitopes with the sequence QPEQPFP and variants thereof in multiple copies, and they typically also harbored many different gluten T-cell epitopes. In the pull-down setting where antibodies were immobilized on a solid phase, peptide fragments with multivalent display of epitopes were targeted. This scenario resembles the situation of the B-cell receptor on the surface of B cells. Conceivably, B cells of celiac disease patients select gluten epitopes that are repeated multiple times in long peptide fragments generated by gut digestive enzymes. As the fragments also contain many different T-cell epitopes, this will lead to generation of strong antibody responses by effective presentation of several distinct T-cell epitopes and establishment of T-cell help to B cells.
Conflict of interest statement
A patent application covering gliadin-reactive monoclonal antibodies has been submitted with Øyvind Steinsbø, Ludvig M. Sollid, Carole H. Dunand and Patrick C. Wilson as inventors.
Figures




Similar articles
-
The preferred substrates for transglutaminase 2 in a complex wheat gluten digest are Peptide fragments harboring celiac disease T-cell epitopes.PLoS One. 2010 Nov 19;5(11):e14056. doi: 10.1371/journal.pone.0014056. PLoS One. 2010. PMID: 21124911 Free PMC article.
-
A quantitative analysis of transglutaminase 2-mediated deamidation of gluten peptides: implications for the T-cell response in celiac disease.J Proteome Res. 2009 Apr;8(4):1748-55. doi: 10.1021/pr800960n. J Proteome Res. 2009. PMID: 19239248
-
Deamidated gliadin peptides form epitopes that transglutaminase antibodies recognize.J Pediatr Gastroenterol Nutr. 2008 Mar;46(3):253-61. doi: 10.1097/MPG.0b013e31815ee555. J Pediatr Gastroenterol Nutr. 2008. PMID: 18376241
-
Coeliac disease--a meeting point for genetics, immunology, and protein chemistry.Lancet. 2003 Apr 12;361(9365):1290-2. doi: 10.1016/s0140-6736(03)12989-3. Lancet. 2003. PMID: 12699968 Review.
-
T-cell and B-cell immunity in celiac disease.Best Pract Res Clin Gastroenterol. 2015 Jun;29(3):413-23. doi: 10.1016/j.bpg.2015.04.001. Epub 2015 May 7. Best Pract Res Clin Gastroenterol. 2015. PMID: 26060106 Review.
Cited by
-
Recent advances in celiac disease and refractory celiac disease.F1000Res. 2019 Jun 26;8:F1000 Faculty Rev-969. doi: 10.12688/f1000research.18701.1. eCollection 2019. F1000Res. 2019. PMID: 31297187 Free PMC article. Review.
-
Coeliac disease: a unique model for investigating broken tolerance in autoimmunity.Clin Transl Immunology. 2016 Nov 2;5(11):e112. doi: 10.1038/cti.2016.58. eCollection 2016 Nov. Clin Transl Immunology. 2016. PMID: 27990287 Free PMC article. Review.
-
Single-cell approaches to dissect adaptive immune responses involved in autoimmunity: the case of celiac disease.Mucosal Immunol. 2022 Jan;15(1):51-63. doi: 10.1038/s41385-021-00452-0. Epub 2021 Sep 16. Mucosal Immunol. 2022. PMID: 34531547 Review.
-
Review Article: Novel Enzyme Therapy Design for Gluten Peptide Digestion Through Exopeptidase Supplementation.Aliment Pharmacol Ther. 2025 Apr;61(7):1123-1139. doi: 10.1111/apt.70014. Epub 2025 Feb 16. Aliment Pharmacol Ther. 2025. PMID: 39955716 Free PMC article. Review.
-
Ancestral Wheat Types Release Fewer Celiac Disease Related T Cell Epitopes than Common Wheat upon Ex Vivo Human Gastrointestinal Digestion.Foods. 2020 Aug 25;9(9):1173. doi: 10.3390/foods9091173. Foods. 2020. PMID: 32854283 Free PMC article.
References
-
- Jabri B. & Sollid L. M. Tissue-mediated control of immunopathology in coeliac disease. Nat.Rev.Immunol. 9, 858–870 (2009). - PubMed
-
- Brandtzaeg P. The changing immunological paradigm in coeliac disease. Immunol.Lett. 105, 127–139 (2006). - PubMed
-
- Douglas A. P., Crabbe P. A. & Hobbs J. R. Immunochemical studies on the serum, intestinal secretions and intestinal mucosa in patients with adult celiac disease and other forms of the celiac syndrome. Gastroenterology 59, 414–425 (1970). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

