Arbuscular mycorrhiza Symbiosis Induces a Major Transcriptional Reprogramming of the Potato SWEET Sugar Transporter Family
- PMID: 27148312
- PMCID: PMC4830831
- DOI: 10.3389/fpls.2016.00487
Arbuscular mycorrhiza Symbiosis Induces a Major Transcriptional Reprogramming of the Potato SWEET Sugar Transporter Family
Abstract
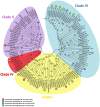
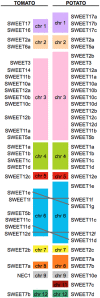
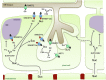
Biotrophic microbes feeding on plants must obtain carbon from their hosts without killing the cells. The symbiotic Arbuscular mycorrhizal (AM) fungi colonizing plant roots do so by inducing major transcriptional changes in the host that ultimately also reprogram the whole carbon partitioning of the plant. AM fungi obtain carbohydrates from the root cortex apoplast, in particular from the periarbuscular space that surrounds arbuscules. However, the mechanisms by which cortical cells export sugars into the apoplast for fungal nutrition are unknown. Recently a novel type of sugar transporter, the SWEET, able to perform not only uptake but also efflux from cells was identified. Plant SWEETs have been shown to be involved in the feeding of pathogenic microbes and are, therefore, good candidates to play a similar role in symbiotic associations. Here we have carried out the first phylogenetic and expression analyses of the potato SWEET family and investigated its role during mycorrhiza symbiosis. The potato genome contains 35 SWEETs that cluster into the same four clades defined in Arabidopsis. Colonization of potato roots by the AM fungus Rhizophagus irregularis imposes major transcriptional rewiring of the SWEET family involving, only in roots, changes in 22 of the 35 members. None of the SWEETs showed mycorrhiza-exclusive induction and most of the 12 induced genes belong to the putative hexose transporters of clade I and II, while only two are putative sucrose transporters from clade III. In contrast, most of the repressed transcripts (10) corresponded to clade III SWEETs. Promoter-reporter assays for three of the induced genes, each from one cluster, showed re-localization of expression to arbuscule-containing cells, supporting a role for SWEETs in the supply of sugars at biotrophic interfaces. The complex transcriptional regulation of SWEETs in roots in response to AM fungal colonization supports a model in which symplastic sucrose in cortical cells could be cleaved in the cytoplasm by sucrose synthases or cytoplasmic invertases and effluxed as glucose, but also directly exported as sucrose and then converted into glucose and fructose by cell wall-bound invertases. Precise biochemical, physiological and molecular analyses are now required to profile the role of each potato SWEET in the arbuscular mycorrhizal symbiosis.
Keywords: Arbuscular mycorrhiza; SWEET transporters; plants; potato; root; sugar transport; symbiosis.
Figures



Similar articles
-
Overexpression of the Potato Monosaccharide Transporter StSWEET7a Promotes Root Colonization by Symbiotic and Pathogenic Fungi by Increasing Root Sink Strength.Front Plant Sci. 2022 Mar 24;13:837231. doi: 10.3389/fpls.2022.837231. eCollection 2022. Front Plant Sci. 2022. PMID: 35401641 Free PMC article.
-
The soybean sugar transporter GmSWEET6 participates in sucrose transport towards fungi during arbuscular mycorrhizal symbiosis.Plant Cell Environ. 2024 Apr;47(4):1041-1052. doi: 10.1111/pce.14772. Epub 2023 Nov 23. Plant Cell Environ. 2024. PMID: 37997205
-
A Eucalyptus Pht1 Family Gene EgPT8 Is Essential for Arbuscule Elongation of Rhizophagus irregularis.Microbiol Spectr. 2022 Dec 21;10(6):e0147022. doi: 10.1128/spectrum.01470-22. Epub 2022 Oct 13. Microbiol Spectr. 2022. PMID: 36227088 Free PMC article.
-
Arbuscular mycorrhiza: biological, chemical, and molecular aspects.J Chem Ecol. 2003 Sep;29(9):1955-79. doi: 10.1023/a:1025695032113. J Chem Ecol. 2003. PMID: 14584670 Review.
-
Multifarious and Interactive Roles of GRAS Transcription Factors During Arbuscular Mycorrhiza Development.Front Plant Sci. 2022 Mar 28;13:836213. doi: 10.3389/fpls.2022.836213. eCollection 2022. Front Plant Sci. 2022. PMID: 35419017 Free PMC article. Review.
Cited by
-
Overexpression of the Potato Monosaccharide Transporter StSWEET7a Promotes Root Colonization by Symbiotic and Pathogenic Fungi by Increasing Root Sink Strength.Front Plant Sci. 2022 Mar 24;13:837231. doi: 10.3389/fpls.2022.837231. eCollection 2022. Front Plant Sci. 2022. PMID: 35401641 Free PMC article.
-
Sugar Partitioning between Ustilago maydis and Its Host Zea mays L during Infection.Plant Physiol. 2019 Apr;179(4):1373-1385. doi: 10.1104/pp.18.01435. Epub 2018 Dec 28. Plant Physiol. 2019. PMID: 30593452 Free PMC article.
-
A coordinated switch in sucrose and callose metabolism enables enhanced symplastic unloading in potato tubers.Quant Plant Biol. 2024 Apr 17;5:e4. doi: 10.1017/qpb.2024.4. eCollection 2024. Quant Plant Biol. 2024. PMID: 38689753 Free PMC article.
-
Cross-kingdom nutrient exchange in the plant-arbuscular mycorrhizal fungus-bacterium continuum.Nat Rev Microbiol. 2024 Dec;22(12):773-790. doi: 10.1038/s41579-024-01073-7. Epub 2024 Jul 16. Nat Rev Microbiol. 2024. PMID: 39014094 Review.
-
The SWEET14 sugar transporter mediates mycorrhizal symbiosis and carbon allocation in Dendrobium officinale.BMC Plant Biol. 2025 Apr 2;25(1):416. doi: 10.1186/s12870-025-06443-8. BMC Plant Biol. 2025. PMID: 40175892 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

