Differential Neuroproteomic and Systems Biology Analysis of Spinal Cord Injury
- PMID: 27150525
- PMCID: PMC4937511
- DOI: 10.1074/mcp.M116.058115
Differential Neuroproteomic and Systems Biology Analysis of Spinal Cord Injury
Abstract
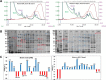
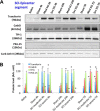
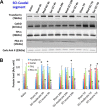
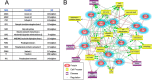
Acute spinal cord injury (SCI) is a devastating condition with many consequences and no known effective treatment. Although it is quite easy to diagnose traumatic SCI, the assessment of injury severity and projection of disease progression or recovery are often challenging, as no consensus biomarkers have been clearly identified. Here rats were subjected to experimental moderate or severe thoracic SCI. At 24h and 7d postinjury, spinal cord segment caudal to injury center versus sham samples was harvested and subjected to differential proteomic analysis. Cationic/anionic-exchange chromatography, followed by 1D polyacrylamide gel electrophoresis, was used to reduce protein complexity. A reverse phase liquid chromatography-tandem mass spectrometry proteomic platform was then utilized to identify proteome changes associated with SCI. Twenty-two and 22 proteins were up-regulated at 24 h and 7 day after SCI, respectively; whereas 19 and 16 proteins are down-regulated at 24 h and 7 day after SCI, respectively, when compared with sham control. A subset of 12 proteins were identified as candidate SCI biomarkers - TF (Transferrin), FASN (Fatty acid synthase), NME1 (Nucleoside diphosphate kinase 1), STMN1 (Stathmin 1), EEF2 (Eukaryotic translation elongation factor 2), CTSD (Cathepsin D), ANXA1 (Annexin A1), ANXA2 (Annexin A2), PGM1 (Phosphoglucomutase 1), PEA15 (Phosphoprotein enriched in astrocytes 15), GOT2 (Glutamic-oxaloacetic transaminase 2), and TPI-1 (Triosephosphate isomerase 1), data are available via ProteomeXchange with identifier PXD003473. In addition, Transferrin, Cathepsin D, and TPI-1 and PEA15 were further verified in rat spinal cord tissue and/or CSF samples after SCI and in human CSF samples from moderate/severe SCI patients. Lastly, a systems biology approach was utilized to determine the critical biochemical pathways and interactome in the pathogenesis of SCI. Thus, SCI candidate biomarkers identified can be used to correlate with disease progression or to identify potential SCI therapeutic targets.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures







References
-
- Cortez R., and Levi A. D. (2008) Acute spinal cord injury. Curr. Treat Options Neurol. 9, 115–125 - PubMed
-
- The Christopher & Dana Reeve Foundation (2008) Paralysis 2008, 1–48
-
- Baptiste D. C., and Fehlings M. G. (2006) Pharmacological approaches to repair the injured spinal cord. J. Neurotrauma 23, 318–334 - PubMed
-
- Yu C.-G., and Geddes J. W. (2007) Sustained calpain inhibition improves locomotor function and tissue sparing following contusive spinal cord injury. Neurochem. Res. 32, 2046–2053 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

