Functional role of TRIM E3 ligase oligomerization and regulation of catalytic activity
- PMID: 27154206
- PMCID: PMC4864278
- DOI: 10.15252/embj.201593741
Functional role of TRIM E3 ligase oligomerization and regulation of catalytic activity
Abstract
TRIM E3 ubiquitin ligases regulate a wide variety of cellular processes and are particularly important during innate immune signalling events. They are characterized by a conserved tripartite motif in their N-terminal portion which comprises a canonical RING domain, one or two B-box domains and a coiled-coil region that mediates ligase dimerization. Self-association via the coiled-coil has been suggested to be crucial for catalytic activity of TRIMs; however, the precise molecular mechanism underlying this observation remains elusive. Here, we provide a detailed characterization of the TRIM ligases TRIM25 and TRIM32 and show how their oligomeric state is linked to catalytic activity. The crystal structure of a complex between the TRIM25 RING domain and an ubiquitin-loaded E2 identifies the structural and mechanistic features that promote a closed E2~Ub conformation to activate the thioester for ubiquitin transfer allowing us to propose a model for the regulation of activity in the full-length protein. Our data reveal an unexpected diversity in the self-association mechanism of TRIMs that might be crucial for their biological function.
Keywords: TRIM25; TRIM32; enzyme mechanism; protein structure; ubiquitin ligase.
© 2016 Francis Crick Institute. Published under the terms of the CC BY 4.0 license.
Figures

- A
Domain organization of
TRIM 25 andTRIM 32 including fragments used in this study. - B, C
(B)
SEC ‐MALLS traces of differentTRIM 25 and (C)TRIM 32 constructs. The proteins are colour coded, and the domain architecture is reported next to the respectiveMALLS curve. The constructs were analysed over different concentration ranges, and the data are reported for: T25R, 4 mg/ml; T25RB 1, 3.2 mg/ml; T25RB 1B2, 3.6 mg/ml; T25RBCC , 0.5 mg/ml; T32R (core and extended), 3 mg/ml; T32RB , 4 mg/ml; T32RBCC, 4 mg/ml.

- A–D
(A, C)
UBE 2D3˜Ub discharge assays with differentTRIM 25 andTRIM 32 constructs, respectively. Assays were carried out withTRIM constructs as indicated and the reaction was monitored over 30 min. The asterisk indicates the band for theTRIM construct. (B, D) Quantification of the discharge assays usingUBE 2D1˜UbAtto andTRIM 25 orTRIM 32, respectively. The loss ofUBE 2D1˜UbAtto is plotted as the average of experimental triplicates (± s.d.). - E–H
(E, G) K63 poly‐ubiquitination assays using
UBE 2N/UBE 2V1 andTRIM 25 andTRIM 32, respectively. Reactions were incubated for 30 min and samples taken at the indicated times. The asterisk indicates the band for theTRIM construct. (F, H) Assays were quantified (see Materials and Methods) by supplementing the reaction with 1 μM UbAtto and integrating the loss of free UbAtto forTRIM 25 andTRIM 32, respectively. Assays were carried out in triplicate (± s.d.).
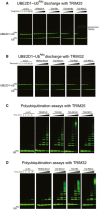
Representative gels of the discharge assays with
UBE 2D1˜UbAtto and differentTRIM 25 constructs. The gels were scanned with a Storm 869 Scanner and the bands for free UbAtto integrated.Representative gels of the discharge assays with
UBE 2D1˜UbAtto and differentTRIM 32 constructs. The gels were scanned with a Storm 869 Scanner and the bands for free UbAtto integrated.Poly‐ubiquitination assays using
UBE 2N/UBE 2V1 and differentTRIM 25 constructs supplemented with fluorescent UbAtto.Poly‐ubiquitination assays using
UBE 2N/UBE 2V1 and differentTRIM 32 constructs supplemented with fluorescent UbAtto.

Structure of the
TRIM 32RING dimer in ribbon representation with eachRING monomer coloured in cyan and blue and the Zn2+ ions as grey spheres.Overlap of the
RING dimers ofTRIM 32 (cyan, 5FEY .pdb),TRIM 25 (yellow, 5FER .pdb) andTRIM 5α (orange, 4TKP .pdb). The structures were overlapped on the circledRING domain. This overlap shows that the structures of theRING s are very similar but that there are differences in the relative orientations of the twoRING s.Structure of the
TRIM 25RING /E2˜Ub complex with theRING domains in the same colour scheme as in (A),UBE 2D1 in grey and the ubiquitin molecules in salmon and red.

Close‐up of the
TRIM 25RING dimer interface highlighting the hydrophobic interactions made between the four α‐helices (left). Close‐up of the interface between eachRING monomer and the proximal ubiquitin (right).UBE 2D1˜UbAtto discharge assays withTRIM 25 wild‐typeRING , the fusedRING constructs and different mutants important for dimerization or the interaction with ubiquitin. Time point zero for the T25‐R Linker and T25‐R Linker V72R samples was taken before the addition of E3 as discharge is very fast. The loss ofUBE 2D1˜UbAtto is plotted as the average of experimental duplicates (± s.d.).UBE 2D3˜Ub discharge assays with the same mutants as in (B), stained with InstantBlue.K63 poly‐ubiquitination assays using
UBE 2N/UBE 2V1. The asterisk indicates the band for theTRIM construct.

Representative gels of the discharge assays with
UBE 2D1˜UbAtto and differentTRIM 25RING mutants. The gels were scanned with a Storm 869 Scanner and the bands for free UbAtto integrated.Representative gels for the equivalent assays with
TRIM 32RING mutants.

- A
Role of E10 in the
RING ofTRIM 25 (blue) in stabilizing the closed E2˜Ub conformation by contacting K11 from the proximal ubiquitin (salmon) and N71 of the oppositeRING (cyan) which in turn contacts K33 of the same ubiquitin. - B
E2˜Ub discharge and K63 poly‐ubiquitination assays to test the role of E9 and E10 in
TRIM 25 activity. Substitution of Glu9 with Arg has no significant effect on activity, whereas the E10R mutation almost completely abolishes catalytic activity. - C
Discharge and K63 poly‐ubiquitination assays to test the role of the equivalent residue E16 in
TRIM 32 and the role of I85R. Mutation of E16R abolishes catalytic activity indicating that the role of the glutamate is conserved. - D, E
Quantification of
UBE 2D1˜UbAtto discharge assays. The loss of E2˜Ub is plotted as the average of experimental duplicates (± s.d.).
- A–D
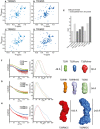
2D 1H‐15N‐
HSQC spectra of theTRIM 25RING domain at a concentration of 125 μM (A) and 570 μM (B), the monomericTRIM 32RING core (C) andTRIM 32RING dimer (D). - E
Isotropic rotational correlation times of the different
TRIM 25 andTRIM 32RING constructs, obtained from the relaxation analysis of resonances in the corresponding 2D 1H‐15NHSQC spectra (dark grey) or calculated from available structures byHYDRONMR (light grey). Error bars are derived from relaxation rate analysis implemented in TENSOR2 (Dosset et al, 2000). - F
Scattering profiles of
RING constructs (left) and their normalized pair‐distribution functions P(r) (middle). The right‐hand panel shows low‐resolution ab initio models derived from theSAXS data analysis. - G
Scattering profiles, their normalized pair‐distribution functions P(r) and low‐resolution ab initio models for
RING and B‐box‐containing constructs. - H
Scattering profiles, their normalized pair‐distribution functions P(r) and low‐resolution ab initio models for the
RBCC domains ofTRIM 25 andTRIM 32. The curves and envelopes are reported using the same colour scheme in all respective panels and the envelopes for the constructs are drawn to scale.

- A, B
Size‐exclusion chromatography in line with
SAXS data recording forTRIM 25RBCC (A) andTRIM 32RBCC (B) at theSWING beamline atSOLEIL . The intensity (blue and green) and Rg (red and purple) profiles are reported as a function of the frames recorded at equal time intervals. - C
Ab initio low‐resolution envelopes calculated with the program
DAMAVER forTRIM 25RING (green),TRIM 32RING core (blue) andTRIM 32RING dimer superimposed to their structures by the programSUPCOMB . The values of the chi‐square are calculated by the programCRYSOL . - D
Low‐resolution envelopes calculated by
DAMAVER forTRIM 25RBCC (red) andTRIM 32RBCC (blue) in different orientations presented in scale with the structures ofTRIM 5αBCC (PDB 4TN 3),TRIM 25CC (PDB 4LTB ) andTRIM 20CC (PDB 4CG 4).


- A, B
The internal volume of the low‐resolution average envelopes of
TRIM 25 (A) andTRIM 32 (B)RBCC , obtained from theSAXS analysis, can accommodate the two monomericTRIM 25RB 1B2 envelopes (represented in blue, see Fig 6) and the two dimericTRIM 32RB 2 envelopes (in green, see Fig 6) at the N‐terminus on each side of the central coiled‐coil spanning the length of the molecule. The structure coordinates ofTRIM 25CC andTRIM 20CC were used to model the central coiled‐coil region ofTRIM 25RBCC andTRIM 32RBCC , respectively. ForTRIM 32RBCC , 2 copies of the same coiled‐coil coordinates (rotated of 180° relative to the log helical axis) were used.
Similar articles
-
Structure and catalytic activation of the TRIM23 RING E3 ubiquitin ligase.Proteins. 2017 Oct;85(10):1957-1961. doi: 10.1002/prot.25348. Epub 2017 Jul 24. Proteins. 2017. PMID: 28681414 Free PMC article.
-
Analysis of the Zn-Binding Domains of TRIM32, the E3 Ubiquitin Ligase Mutated in Limb Girdle Muscular Dystrophy 2H.Cells. 2019 Mar 16;8(3):254. doi: 10.3390/cells8030254. Cells. 2019. PMID: 30884854 Free PMC article.
-
Does it take two to tango? RING domain self-association and activity in TRIM E3 ubiquitin ligases.Biochem Soc Trans. 2020 Dec 18;48(6):2615-2624. doi: 10.1042/BST20200383. Biochem Soc Trans. 2020. PMID: 33170204 Free PMC article. Review.
-
Tripartite motif ligases catalyze polyubiquitin chain formation through a cooperative allosteric mechanism.J Biol Chem. 2013 Mar 22;288(12):8209-8221. doi: 10.1074/jbc.M113.451567. Epub 2013 Feb 13. J Biol Chem. 2013. PMID: 23408431 Free PMC article.
-
TRIM E3 Ubiquitin Ligases in Rare Genetic Disorders.Adv Exp Med Biol. 2020;1233:311-325. doi: 10.1007/978-3-030-38266-7_14. Adv Exp Med Biol. 2020. PMID: 32274764 Review.
Cited by
-
Mapping the interaction between Trim28 and the KRAB domain at the center of Trim28 silencing of endogenous retroviruses.Protein Sci. 2022 Oct;31(10):e4436. doi: 10.1002/pro.4436. Protein Sci. 2022. PMID: 36173157 Free PMC article.
-
A mechanism in agrin signaling revealed by a prevalent Rapsyn mutation in congenital myasthenic syndrome.Elife. 2019 Sep 24;8:e49180. doi: 10.7554/eLife.49180. Elife. 2019. PMID: 31549961 Free PMC article.
-
TRIM17 and TRIM28 antagonistically regulate the ubiquitination and anti-apoptotic activity of BCL2A1.Cell Death Differ. 2019 May;26(5):902-917. doi: 10.1038/s41418-018-0169-5. Epub 2018 Jul 24. Cell Death Differ. 2019. PMID: 30042493 Free PMC article.
-
General Model for Retroviral Capsid Pattern Recognition by TRIM5 Proteins.J Virol. 2018 Jan 30;92(4):e01563-17. doi: 10.1128/JVI.01563-17. Print 2018 Feb 15. J Virol. 2018. PMID: 29187540 Free PMC article.
-
Ubiquitin transfer by a RING E3 ligase occurs from a closed E2~ubiquitin conformation.Nat Commun. 2020 Jun 5;11(1):2846. doi: 10.1038/s41467-020-16666-y. Nat Commun. 2020. PMID: 32503993 Free PMC article.
References
-
- Adams PD, Afonine PV, Bunkoczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung LW, Kapral GJ, Grosse‐Kunstleve RW, McCoy AJ, Moriarty NW, Oeffner R, Read RJ, Richardson DC, Richardson JS, Terwilliger TC, Zwart PH (2010) PHENIX: a comprehensive Python‐based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr 66: 213–221 - PMC - PubMed
-
- Berndsen CE, Wolberger C (2014) New insights into ubiquitin E3 ligase mechanism. Nat Struct Mol Biol 21: 301–307 - PubMed
-
- Brzovic PS, Rajagopal P, Hoyt DW, King MC, Klevit RE (2001) Structure of a BRCA1‐BARD1 heterodimeric RING‐RING complex. Nat Struct Biol 8: 833–837 - PubMed
-
- Carvalho AF, Pinto MP, Grou CP, Vitorino R, Domingues P, Yamao F, Sa‐Miranda C, Azevedo JE (2012) High‐yield expression in Escherichia coli and purification of mouse ubiquitin‐activating enzyme E1. Mol Biotechnol 51: 254–261 - PubMed
-
- Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6: 277–293 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

