Transcriptional signatures of brain aging and Alzheimer's disease: What are our rodent models telling us?
- PMID: 27155503
- PMCID: PMC5533609
- DOI: 10.1016/j.bbr.2016.05.007
Transcriptional signatures of brain aging and Alzheimer's disease: What are our rodent models telling us?
Abstract
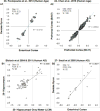
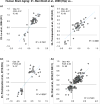
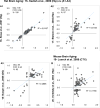
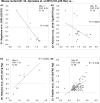
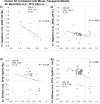
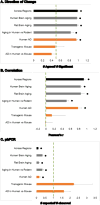
Aging is the biggest risk factor for idiopathic Alzheimer's disease (AD). Recently, the National Institutes of Health released AD research recommendations that include: appreciating normal brain aging, expanding data-driven research, using open-access resources, and evaluating experimental reproducibility. Transcriptome data sets for aging and AD in humans and animal models are available in NIH-curated, publically accessible databases. However, little work has been done to test for concordance among those molecular signatures. Here, we test the hypothesis that brain transcriptional profiles from animal models recapitulate those observed in the human condition. Raw transcriptional profile data from twenty-nine studies were analyzed to produce p-values and fold changes for young vs. aged or control vs. AD conditions. Concordance across profiles was assessed at three levels: (1) # of significant genes observed vs. # expected by chance; (2) proportion of significant genes showing directional agreement; (3) correlation among studies for magnitude of effect among significant genes. The highest concordance was found within subjects across brain regions. Normal brain aging was concordant across studies, brain regions, and species, despite profound differences in chronological aging among humans, rats and mice. Human studies of idiopathic AD were concordant across brain structures and studies, but were not concordant with the transcriptional profiles of transgenic AD mouse models. Further, the five transgenic AD mouse models that were assessed were not concordant with one another. These results suggest that normal brain aging is similar in humans and research animals, and that different transgenic AD model mice may reflect selected aspects of AD pathology.
Keywords: Aging; Alzheimer’s disease; Transcriptome.
Copyright © 2016 The Authors. Published by Elsevier B.V. All rights reserved.
Figures







Similar articles
-
Comparative analysis of cortical gene expression in mouse models of Alzheimer's disease.Neurobiol Aging. 2006 Mar;27(3):377-86. doi: 10.1016/j.neurobiolaging.2005.02.010. Epub 2005 May 31. Neurobiol Aging. 2006. PMID: 15927307
-
The Alzheimer's disease transcriptome mimics the neuroprotective signature of IGF-1 receptor-deficient neurons.Brain. 2017 Jul 1;140(7):2012-2027. doi: 10.1093/brain/awx132. Brain. 2017. PMID: 28595357
-
Distinct transcriptome expression of the temporal cortex of the primate Microcebus murinus during brain aging versus Alzheimer's disease-like pathology.PLoS One. 2010 Sep 16;5(9):e12770. doi: 10.1371/journal.pone.0012770. PLoS One. 2010. PMID: 20862281 Free PMC article.
-
A framework to understand the variations of PSD-95 expression in brain aging and in Alzheimer's disease.Ageing Res Rev. 2014 Nov;18:86-94. doi: 10.1016/j.arr.2014.09.004. Epub 2014 Sep 26. Ageing Res Rev. 2014. PMID: 25264360 Review.
-
Rodent models for Alzheimer's disease drug discovery.Expert Opin Drug Discov. 2015 Jul;10(7):703-11. doi: 10.1517/17460441.2015.1041913. Epub 2015 Apr 30. Expert Opin Drug Discov. 2015. PMID: 25927677 Free PMC article. Review.
Cited by
-
Genetic architecture of Alzheimer's disease.Neurobiol Dis. 2020 Sep;143:104976. doi: 10.1016/j.nbd.2020.104976. Epub 2020 Jun 18. Neurobiol Dis. 2020. PMID: 32565066 Free PMC article. Review.
-
Cognitive Reserve in Model Systems for Mechanistic Discovery: The Importance of Longitudinal Studies.Front Aging Neurosci. 2021 Jan 21;12:607685. doi: 10.3389/fnagi.2020.607685. eCollection 2020. Front Aging Neurosci. 2021. PMID: 33551788 Free PMC article. Review.
-
Presenilin-2 and Calcium Handling: Molecules, Organelles, Cells and Brain Networks.Cells. 2020 Sep 25;9(10):2166. doi: 10.3390/cells9102166. Cells. 2020. PMID: 32992716 Free PMC article. Review.
-
Brain transcriptomes of zebrafish and mouse Alzheimer's disease knock-in models imply early disrupted energy metabolism.Dis Model Mech. 2022 Jan 1;15(1):dmm049187. doi: 10.1242/dmm.049187. Epub 2022 Jan 26. Dis Model Mech. 2022. PMID: 34842276 Free PMC article.
-
Dynamic Changes of Endogenic or Exogenic β-Carboline Alkaloid Harmine in Different Mammals and Human in vivo at Developmental and Physiological States.Front Aging Neurosci. 2022 Jan 14;13:773638. doi: 10.3389/fnagi.2021.773638. eCollection 2021. Front Aging Neurosci. 2022. PMID: 35095466 Free PMC article.
References
-
- Lister JP, Barnes CA. Neurobiological changes in the hippocampus during normative aging. Arch Neurol. 2009;66:829–33. - PubMed
-
- Hebert LE, Scherr PA, Bienias JL, Bennett DA, Evans DA. Alzheimer disease in the US population: prevalence estimates using the 2000 census. Arch Neurol. 2003;60:1119–22. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

