Stop pulling my strings - what telomeres taught us about the DNA damage response
- PMID: 27165790
- PMCID: PMC5385261
- DOI: 10.1038/nrm.2016.43
Stop pulling my strings - what telomeres taught us about the DNA damage response
Abstract
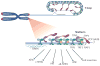
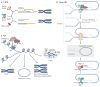
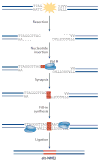
Mammalian cells have evolved specialized mechanisms to sense and repair double-strand breaks (DSBs) to maintain genomic stability. However, in certain cases, the activity of these pathways can lead to aberrant DNA repair, genomic instability and tumorigenesis. One such case is DNA repair at the natural ends of linear chromosomes, known as telomeres, which can lead to chromosome-end fusions. Here, we review data obtained over the past decade and discuss the mechanisms that protect mammalian chromosome ends from the DNA damage response. We also discuss how telomere research has helped to uncover key steps in DSB repair. Last, we summarize how dysfunctional telomeres and the ensuing genomic instability drive the progression of cancer.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Telomeres and double-strand breaks - all's well that "ends" well. ..Radiat Res. 2008 Jan;169(1):1-7. doi: 10.1667/RR1197.1. Radiat Res. 2008. PMID: 18159961
-
Telomere dysfunction and genome instability.Front Biosci (Landmark Ed). 2012 Jun 1;17(6):2181-96. doi: 10.2741/4044. Front Biosci (Landmark Ed). 2012. PMID: 22652771 Review.
-
The role of the nonhomologous end-joining DNA double-strand break repair pathway in telomere biology.Annu Rev Genet. 2006;40:237-77. doi: 10.1146/annurev.genet.39.110304.095755. Annu Rev Genet. 2006. PMID: 16822175 Review.
-
The DNA damage response at dysfunctional telomeres, and at interstitial and subtelomeric DNA double-strand breaks.Genes Genet Syst. 2018 Jan 20;92(3):135-152. doi: 10.1266/ggs.17-00014. Epub 2017 Nov 22. Genes Genet Syst. 2018. PMID: 29162774 Review.
-
Mechanisms and regulation of DNA end resection.EMBO J. 2010 Sep 1;29(17):2864-74. doi: 10.1038/emboj.2010.165. Epub 2010 Jul 20. EMBO J. 2010. PMID: 20647996 Free PMC article. Review.
Cited by
-
Fanconi anemia and dyskeratosis congenita/telomere biology disorders: Two inherited bone marrow failure syndromes with genomic instability.Front Oncol. 2022 Aug 25;12:949435. doi: 10.3389/fonc.2022.949435. eCollection 2022. Front Oncol. 2022. PMID: 36091172 Free PMC article. Review.
-
A non-catalytic N-terminus domain of WRN prevents mitotic telomere deprotection.Sci Rep. 2023 Jan 12;13(1):645. doi: 10.1038/s41598-023-27598-0. Sci Rep. 2023. PMID: 36635307 Free PMC article.
-
The dynamics of forming a triplex in an artificial telomere inferred by DNA mechanics.Nucleic Acids Res. 2019 Sep 5;47(15):e86. doi: 10.1093/nar/gkz464. Nucleic Acids Res. 2019. PMID: 31114915 Free PMC article.
-
SERPINA3 is a key modulator of HNRNP-K transcriptional activity against oxidative stress in HCC.Redox Biol. 2019 Jun;24:101217. doi: 10.1016/j.redox.2019.101217. Epub 2019 May 12. Redox Biol. 2019. PMID: 31121493 Free PMC article.
-
High-Risk Alphapapillomavirus Oncogenes Impair the Homologous Recombination Pathway.J Virol. 2017 Sep 27;91(20):e01084-17. doi: 10.1128/JVI.01084-17. Print 2017 Oct 15. J Virol. 2017. PMID: 28768872 Free PMC article.
References
-
- Greider CW. Telomerase and telomere-length regulation: lessons from small eukaryotes to mammals. Cold Spring Harb Symp Quant Biol. 1993;58:719–723. - PubMed
-
- Makarov VL, Hirose Y, Langmore JP. Long G tails at both ends of human chromosomes suggest a C strand degradation mechanism for telomere shortening. Cell. 1997;88:657–666. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

