Quantitative Mass Spectrometry Reveals Changes in Histone H2B Variants as Cells Undergo Inorganic Arsenic-Mediated Cellular Transformation
- PMID: 27169413
- PMCID: PMC4937513
- DOI: 10.1074/mcp.M116.058412
Quantitative Mass Spectrometry Reveals Changes in Histone H2B Variants as Cells Undergo Inorganic Arsenic-Mediated Cellular Transformation
Abstract
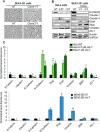
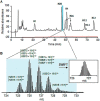
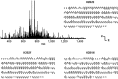
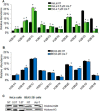
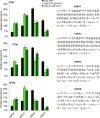
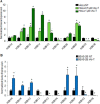
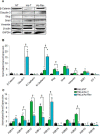
Exposure to inorganic arsenic, a ubiquitous environmental toxic metalloid, leads to carcinogenesis. However, the mechanism is unknown. Several studies have shown that inorganic arsenic exposure alters specific gene expression patterns, possibly through alterations in chromatin structure. While most studies on understanding the mechanism of chromatin-mediated gene regulation have focused on histone post-translational modifications, the role of histone variants remains largely unknown. Incorporation of histone variants alters the functional properties of chromatin. To understand the global dynamics of chromatin structure and function in arsenic-mediated carcinogenesis, analysis of the histone variants incorporated into the nucleosome and their covalent modifications is required. Here we report the first global mass spectrometric analysis of histone H2B variants as cells undergo arsenic-mediated epithelial to mesenchymal transition. We used electron capture dissociation-based top-down tandem mass spectrometry analysis validated with quantitative reverse transcription real-time polymerase chain reaction to identify changes in the expression levels of H2B variants in inorganic arsenic-mediated epithelial-mesenchymal transition. We identified changes in the expression levels of specific histone H2B variants in two cell types, which are dependent on dose and length of exposure of inorganic arsenic. In particular, we found increases in H2B variants H2B1H/1K/1C/1J/1O and H2B2E/2F, and significant decreases in H2B1N/1D/1B as cells undergo inorganic arsenic-mediated epithelial-mesenchymal transition. The analysis of these histone variants provides a first step toward an understanding of the functional significance of the diversity of histone structures, especially in inorganic arsenic-mediated gene expression and carcinogenesis.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures
Similar articles
-
Transient and permanent changes in DNA methylation patterns in inorganic arsenic-mediated epithelial-to-mesenchymal transition.Toxicol Appl Pharmacol. 2017 Sep 15;331:6-17. doi: 10.1016/j.taap.2017.03.017. Epub 2017 Mar 21. Toxicol Appl Pharmacol. 2017. PMID: 28336213 Free PMC article.
-
Characterization of histone H2A and H2B variants and their post-translational modifications by mass spectrometry.Mol Cell Proteomics. 2006 Mar;5(3):541-52. doi: 10.1074/mcp.M500288-MCP200. Epub 2005 Nov 30. Mol Cell Proteomics. 2006. PMID: 16319397
-
Inorganic Arsenic-induced cellular transformation is coupled with genome wide changes in chromatin structure, transcriptome and splicing patterns.BMC Genomics. 2015 Mar 19;16(1):212. doi: 10.1186/s12864-015-1295-9. BMC Genomics. 2015. PMID: 25879800 Free PMC article.
-
The ABCs of the H2Bs: The histone H2B sequences, variants, and modifications.Trends Genet. 2025 Jun;41(6):506-521. doi: 10.1016/j.tig.2025.01.003. Epub 2025 Feb 20. Trends Genet. 2025. PMID: 39984351 Review.
-
Histone variants: key players of chromatin.Cell Tissue Res. 2014 Jun;356(3):457-66. doi: 10.1007/s00441-014-1862-4. Epub 2014 Apr 30. Cell Tissue Res. 2014. PMID: 24781148 Review.
Cited by
-
Top-Down Proteomics: Ready for Prime Time?Anal Chem. 2018 Jan 2;90(1):110-127. doi: 10.1021/acs.analchem.7b04747. Epub 2017 Dec 15. Anal Chem. 2018. PMID: 29161012 Free PMC article. Review.
-
Metals and Mechanisms of Carcinogenesis.Annu Rev Pharmacol Toxicol. 2019 Jan 6;59:537-554. doi: 10.1146/annurev-pharmtox-010818-021031. Annu Rev Pharmacol Toxicol. 2019. PMID: 30625284 Free PMC article. Review.
-
Transient and permanent changes in DNA methylation patterns in inorganic arsenic-mediated epithelial-to-mesenchymal transition.Toxicol Appl Pharmacol. 2017 Sep 15;331:6-17. doi: 10.1016/j.taap.2017.03.017. Epub 2017 Mar 21. Toxicol Appl Pharmacol. 2017. PMID: 28336213 Free PMC article.
-
A dynamic model of inorganic arsenic-induced carcinogenesis reveals an epigenetic mechanism for epithelial-mesenchymal plasticity.Environ Pollut. 2024 Apr 15;347:123586. doi: 10.1016/j.envpol.2024.123586. Epub 2024 Mar 9. Environ Pollut. 2024. PMID: 38467368 Free PMC article.
-
Microarray dataset of transient and permanent DNA methylation changes in HeLa cells undergoing inorganic arsenic-mediated epithelial-to-mesenchymal transition.Data Brief. 2017 May 10;13:6-9. doi: 10.1016/j.dib.2017.05.002. eCollection 2017 Aug. Data Brief. 2017. PMID: 28589171 Free PMC article.
References
-
- Cheng T. F., Choudhuri S., and Muldoon-Jacobs K. (2012) Epigenetic targets of some toxicologically relevant metals: a review of the literature. JAT 32, 643–653 - PubMed
-
- Huang C., Ke Q., Costa M., and Shi X. (2004) Molecular mechanisms of arsenic carcinogenesis. Mol. Cell. Biochem. 255, 57–66 - PubMed
-
- Lynn S., Lai H. T., Gurr J. R., and Jan K. Y. (1997) Arsenite retards DNA break rejoining by inhibiting DNA ligation. Mutagenesis 12, 353–358 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

