Coarse-grained simulation reveals key features of HIV-1 capsid self-assembly
- PMID: 27174390
- PMCID: PMC4869257
- DOI: 10.1038/ncomms11568
Coarse-grained simulation reveals key features of HIV-1 capsid self-assembly
Abstract
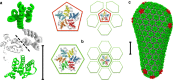
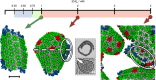
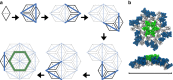
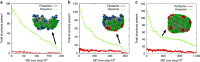
The maturation of HIV-1 viral particles is essential for viral infectivity. During maturation, many copies of the capsid protein (CA) self-assemble into a capsid shell to enclose the viral RNA. The mechanistic details of the initiation and early stages of capsid assembly remain to be delineated. We present coarse-grained simulations of capsid assembly under various conditions, considering not only capsid lattice self-assembly but also the potential disassembly of capsid upon delivery to the cytoplasm of a target cell. The effects of CA concentration, molecular crowding, and the conformational variability of CA are described, with results indicating that capsid nucleation and growth is a multi-stage process requiring well-defined metastable intermediates. Generation of the mature capsid lattice is sensitive to local conditions, with relatively subtle changes in CA concentration and molecular crowding influencing self-assembly and the ensemble of structural morphologies.
Figures



References
-
- Briggs J. A. & Krausslich H. G. The molecular architecture of HIV. J. Mol. Biol. 410, 491–500 (2011). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

