Mitochondrial Polyadenylation Is a One-Step Process Required for mRNA Integrity and tRNA Maturation
- PMID: 27176048
- PMCID: PMC4866704
- DOI: 10.1371/journal.pgen.1006028
Mitochondrial Polyadenylation Is a One-Step Process Required for mRNA Integrity and tRNA Maturation
Abstract
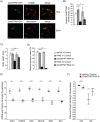
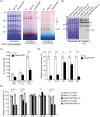
Polyadenylation has well characterised roles in RNA turnover and translation in a variety of biological systems. While polyadenylation on mitochondrial transcripts has been suggested to be a two-step process required to complete translational stop codons, its involvement in mitochondrial RNA turnover is less well understood. We studied knockdown and knockout models of the mitochondrial poly(A) polymerase (MTPAP) in Drosophila melanogaster and demonstrate that polyadenylation of mitochondrial mRNAs is exclusively performed by MTPAP. Further, our results show that mitochondrial polyadenylation does not regulate mRNA stability but protects the 3' terminal integrity, and that despite a lack of functioning 3' ends, these trimmed transcripts are translated, suggesting that polyadenylation is not required for mitochondrial translation. Additionally, loss of MTPAP leads to reduced steady-state levels and disturbed maturation of tRNACys, indicating that polyadenylation in mitochondria might be important for the stability and maturation of specific tRNAs.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




Similar articles
-
The bicoid stability factor controls polyadenylation and expression of specific mitochondrial mRNAs in Drosophila melanogaster.PLoS Genet. 2011 Oct;7(10):e1002324. doi: 10.1371/journal.pgen.1002324. Epub 2011 Oct 13. PLoS Genet. 2011. PMID: 22022283 Free PMC article.
-
Maturation of selected human mitochondrial tRNAs requires deadenylation.Elife. 2017 Jul 26;6:e27596. doi: 10.7554/eLife.27596. Elife. 2017. PMID: 28745585 Free PMC article.
-
Functional polypeptides can be synthesized from human mitochondrial transcripts lacking termination codons.Biochem J. 2004 Feb 1;377(Pt 3):725-31. doi: 10.1042/BJ20031556. Biochem J. 2004. PMID: 14585098 Free PMC article.
-
Mitochondrial poly(A) polymerase and polyadenylation.Biochim Biophys Acta. 2012 Sep-Oct;1819(9-10):992-7. doi: 10.1016/j.bbagrm.2011.10.012. Epub 2011 Dec 7. Biochim Biophys Acta. 2012. PMID: 22172994 Free PMC article. Review.
-
Polyadenylation in mammalian mitochondria: insights from recent studies.Biochim Biophys Acta. 2008 Apr;1779(4):266-9. doi: 10.1016/j.bbagrm.2008.02.001. Epub 2008 Feb 14. Biochim Biophys Acta. 2008. PMID: 18312863 Review.
Cited by
-
The process of mammalian mitochondrial protein synthesis.Cell Tissue Res. 2017 Jan;367(1):5-20. doi: 10.1007/s00441-016-2456-0. Epub 2016 Jul 14. Cell Tissue Res. 2017. PMID: 27411691 Free PMC article. Review.
-
Construction of scFv Antibodies against the Outer Loops of the Microsporidium Nosema bombycis ATP/ADP-Transporters and Selection of the Fragment Efficiently Inhibiting Parasite Growth.Int J Mol Sci. 2022 Dec 4;23(23):15307. doi: 10.3390/ijms232315307. Int J Mol Sci. 2022. PMID: 36499634 Free PMC article.
-
Human Mitochondrial RNA Processing and Modifications: Overview.Int J Mol Sci. 2021 Jul 27;22(15):7999. doi: 10.3390/ijms22157999. Int J Mol Sci. 2021. PMID: 34360765 Free PMC article. Review.
-
Positive Correlation of the Gene Rearrangements and Evolutionary Rates in the Mitochondrial Genomes of Thrips (Insecta: Thysanoptera).Insects. 2022 Jun 27;13(7):585. doi: 10.3390/insects13070585. Insects. 2022. PMID: 35886761 Free PMC article.
-
Simultaneous processing and degradation of mitochondrial RNAs revealed by circularized RNA sequencing.Nucleic Acids Res. 2017 May 19;45(9):5487-5500. doi: 10.1093/nar/gkx104. Nucleic Acids Res. 2017. PMID: 28201688 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

