Bacterial Diversity and Community Structure in Korean Ginseng Field Soil Are Shifted by Cultivation Time
- PMID: 27187071
- PMCID: PMC4871511
- DOI: 10.1371/journal.pone.0155055
Bacterial Diversity and Community Structure in Korean Ginseng Field Soil Are Shifted by Cultivation Time
Abstract
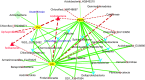
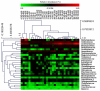
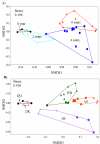
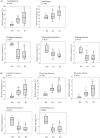
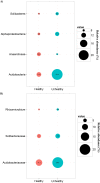
Traditional molecular methods have been used to examine bacterial communities in ginseng-cultivated soil samples in a time-dependent manner. Despite these efforts, our understanding of the bacterial community is still inadequate. Therefore, in this study, a high-throughput sequencing approach was employed to investigate bacterial diversity in various ginseng field soil samples over cultivation times of 2, 4, and 6 years in the first and second rounds of cultivation. We used non-cultivated soil samples to perform a comparative study. Moreover, this study assessed changes in the bacterial community associated with soil depth and the health state of the ginseng. Bacterial richness decreased through years of cultivation. This study detected differences in relative abundance of bacterial populations between the first and second rounds of cultivation, years of cultivation, and health states of ginseng. These bacterial populations were mainly distributed in the classes Acidobacteria, Alphaproteobacteria, Deltaproteobacteria, Gammaproteobacteria, and Sphingobacteria. In addition, we found that pH, available phosphorus, and exchangeable Ca+ seemed to have high correlations with bacterial class in ginseng cultivated soil.
Conflict of interest statement
Figures


Similar articles
-
Diversity and composition of the Panax ginseng rhizosphere microbiome in various cultivation modesand ages.BMC Microbiol. 2021 Jan 8;21(1):18. doi: 10.1186/s12866-020-02081-2. BMC Microbiol. 2021. PMID: 33419388 Free PMC article.
-
Soil pH and electrical conductivity are key edaphic factors shaping bacterial communities of greenhouse soils in Korea.J Microbiol. 2016 Dec;54(12):838-845. doi: 10.1007/s12275-016-6526-5. Epub 2016 Nov 26. J Microbiol. 2016. PMID: 27888456
-
Effects of American Ginseng Cultivation on Bacterial Community Structure and Responses of Soil Nutrients in Different Ecological Niches.J Microbiol Biotechnol. 2022 Apr 28;32(4):419-429. doi: 10.4014/jmb.2202.02003. J Microbiol Biotechnol. 2022. PMID: 35283425 Free PMC article.
-
The Rhizosphere Microbiome of Ginseng.Microorganisms. 2022 Jun 2;10(6):1152. doi: 10.3390/microorganisms10061152. Microorganisms. 2022. PMID: 35744670 Free PMC article. Review.
-
New microorganism isolation techniques with emphasis on laser printing.Int J Bioprint. 2018 Dec 14;5(1):165. doi: 10.18063/ijb.v5i1.165. eCollection 2019. Int J Bioprint. 2018. PMID: 32596530 Free PMC article. Review.
Cited by
-
Ornithinimicrobium panacihumi sp. nov., Antagonistic Bacteria Against Root Rot Fungal Pathogens, Isolated from Cultivated Ginseng Soil.Curr Microbiol. 2019 Jan;76(1):22-28. doi: 10.1007/s00284-018-1579-9. Epub 2018 Oct 31. Curr Microbiol. 2019. PMID: 30382345
-
Different Age-Induced Changes in Rhizosphere Microbial Composition and Function of Panax ginseng in Transplantation Mode.Front Plant Sci. 2020 Nov 12;11:563240. doi: 10.3389/fpls.2020.563240. eCollection 2020. Front Plant Sci. 2020. PMID: 33281838 Free PMC article.
-
Siderophore-producing rhizobacteria reduce heavy metal-induced oxidative stress in Panax ginseng Meyer.J Ginseng Res. 2021 Mar;45(2):218-227. doi: 10.1016/j.jgr.2019.12.008. Epub 2020 Jan 7. J Ginseng Res. 2021. PMID: 33841002 Free PMC article.
-
Microbial diversity and community structure changes in the rhizosphere soils of Atractylodes lancea from different planting years.Plant Signal Behav. 2021 Feb 1;16(2):1854507. doi: 10.1080/15592324.2020.1854507. Epub 2020 Dec 8. Plant Signal Behav. 2021. PMID: 33289592 Free PMC article.
-
Cell-Free Extracts of the Ginseng Soil Bacterium Pseudomonas plecoglossicida Promote Suppression of Resistance of American Ginseng (Panax quinquefolius) to Root Rot Caused by Ilyonectria mors-panacis.Biology (Basel). 2024 Aug 29;13(9):671. doi: 10.3390/biology13090671. Biology (Basel). 2024. PMID: 39336098 Free PMC article.
References
-
- Ernst E (2010) Panax ginseng: An overview of the clinical evidence. J Ginseng Res 34: 259–263.
-
- Bang CS, Hong SH, Suk KT, Kim JB, Han SH, Sung H, et al. (2014) Effects of Korean Red Ginseng (Panax ginseng), urushiol (Rhus vernicifera Stokes), and probiotics (Lactobacillus rhamnosus R0011 and Lactobacillus acidophilus R0052) on the gut–liver axis of alcoholic liver disease. J Ginseng Res 38: 167–172. 10.1016/j.jgr.2014.04.002 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

