AFM-Based Single Molecule Techniques: Unraveling the Amyloid Pathogenic Species
- PMID: 27189600
- PMCID: PMC5080865
- DOI: 10.2174/1381612822666160518141911
AFM-Based Single Molecule Techniques: Unraveling the Amyloid Pathogenic Species
Abstract
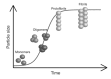
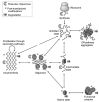
Background: A wide class of human diseases and neurodegenerative disorders, such as Alzheimer's disease, is due to the failure of a specific peptide or protein to keep its native functional conformational state and to undergo a conformational change into a misfolded state, triggering the formation of fibrillar cross-β sheet amyloid aggregates. During the fibrillization, several coexisting species are formed, giving rise to a highly heterogeneous mixture. Despite its fundamental role in biological function and malfunction, the mechanism of protein self-assembly and the fundamental origins of the connection between aggregation, cellular toxicity and the biochemistry of neurodegeneration remains challenging to elucidate in molecular detail. In particular, the nature of the specific state of proteins that is most prone to cause cytotoxicity is not established.
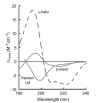
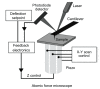
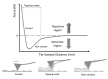
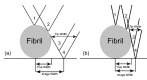
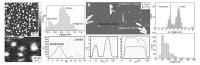
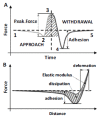
Methods: In the present review, we present the latest advances obtained by Atomic Force Microscopy (AFM) based techniques to unravel the biophysical properties of amyloid aggregates at the nanoscale. Unraveling amyloid single species biophysical properties still represents a formidable experimental challenge, mainly because of their nanoscale dimensions and heterogeneous nature. Bulk techniques, such as circular dichroism or infrared spectroscopy, are not able to characterize the heterogeneity and inner properties of amyloid aggregates at the single species level, preventing a profound investigation of the correlation between the biophysical properties and toxicity of the individual species.
Conclusion: The information delivered by AFM based techniques could be central to study the aggregation pathway of proteins and to design molecules that could interfere with amyloid aggregation delaying the onset of misfolding diseases.
Figures









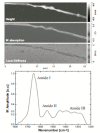

References
-
- Hardy J., Selkoe D.J. The amyloid hypothesis of Alzheimer's disease: progress and problems on the road to therapeutics. Science. 2002;297(5580):353–356. - PubMed
-
- Goedert M. Familial Parkinson's disease. The awakening of alpha-synuclein. Nature. 1997;388(6639):232–233. - PubMed
-
- Zoghbi H.Y., Orr H.T. Glutamine repeats and neurodegeneration. Annu. Rev. Neurosci. 2000;23:217–247. - PubMed
-
- Chiti F., Dobson C.M. Protein misfolding, functional amyloid, and human disease. Annu. Rev. Biochem. 2006;75:333–366. - PubMed
-
- Knowles T.P., Vendruscolo M., Dobson C.M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell Biol. 2014;15(6):384–396. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

