Identification and Characterization of Epstein-Barr Virus Genomes in Lung Carcinoma Biopsy Samples by Next-Generation Sequencing Technology
- PMID: 27189712
- PMCID: PMC4870493
- DOI: 10.1038/srep26156
Identification and Characterization of Epstein-Barr Virus Genomes in Lung Carcinoma Biopsy Samples by Next-Generation Sequencing Technology
Abstract
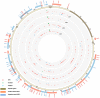
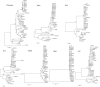
Epstein-Barr virus (EBV) has been detected in the tumor cells of several cancers, including some cases of lung carcinoma (LC). However, the genomic characteristics and diversity of EBV strains associated with LC are poorly understood. In this study, we sequenced the EBV genomes isolated from four primary LC tumor biopsy samples, designated LC1 to LC4. Comparative analysis demonstrated that LC strains were more closely related to GD1 strain. Compared to GD1 reference genome, a total of 520 variations in all, including 498 substitutions, 12 insertions, and 10 deletions were found. Latent genes were found to harbor the most numbers of nonsynonymous mutations. Phylogenetic analysis showed that all LC strains were closely related to Asian EBV strains, whereas different from African/American strains. LC2 genome was distinct from the other three LC genomes, suggesting at least two parental lineages of EBV among the LC genomes may exist. All LC strains could be classified as China 1 and V-val subtype according to the amino acid sequence of LMP1 and EBNA1, respectively. In conclusion, our results showed the genomic diversity among EBV genomes isolated from LC, which might facilitate to uncover the previously unknown variations of pathogenic significance.
Figures



Similar articles
-
Genome-Wide Analysis of Epstein-Barr Virus Isolated from Extranodal NK/T-Cell Lymphoma, Nasal Type.Oncologist. 2019 Sep;24(9):e905-e913. doi: 10.1634/theoncologist.2017-0588. Epub 2019 Apr 2. Oncologist. 2019. PMID: 30940744 Free PMC article.
-
Genome-Wide Analysis of 18 Epstein-Barr Viruses Isolated from Primary Nasopharyngeal Carcinoma Biopsy Specimens.J Virol. 2017 Aug 10;91(17):e00301-17. doi: 10.1128/JVI.00301-17. Print 2017 Sep 1. J Virol. 2017. PMID: 28637758 Free PMC article.
-
Genomic diversity of Epstein-Barr virus genomes isolated from primary nasopharyngeal carcinoma biopsy samples.J Virol. 2014 Sep;88(18):10662-72. doi: 10.1128/JVI.01665-14. Epub 2014 Jul 2. J Virol. 2014. PMID: 24991008 Free PMC article.
-
From Conventional to Next Generation Sequencing of Epstein-Barr Virus Genomes.Viruses. 2016 Feb 24;8(3):60. doi: 10.3390/v8030060. Viruses. 2016. PMID: 26927157 Free PMC article. Review.
-
Phylogenetic comparison of Epstein-Barr virus genomes.J Microbiol. 2018 Aug;56(8):525-533. doi: 10.1007/s12275-018-8039-x. Epub 2018 Jun 14. J Microbiol. 2018. PMID: 29948828 Review.
Cited by
-
Bronchoalveolar Lavage Proteomics in Patients with Suspected Lung Cancer.Sci Rep. 2017 Feb 7;7:42190. doi: 10.1038/srep42190. Sci Rep. 2017. PMID: 28169345 Free PMC article.
-
Screening for viruses in lung adenocarcinoma in China.Chin Med J Pulm Crit Care Med. 2024 Sep 19;2(3):197-199. doi: 10.1016/j.pccm.2024.05.001. eCollection 2024 Sep. Chin Med J Pulm Crit Care Med. 2024. PMID: 39403412 Free PMC article. No abstract available.
-
Detection of Epstein-Barr Virus Infection in Non-Small Cell Lung Cancer.Cancers (Basel). 2019 May 31;11(6):759. doi: 10.3390/cancers11060759. Cancers (Basel). 2019. PMID: 31159203 Free PMC article.
-
Genomic variations in EBNA3C of EBV associate with posttransplant lymphoproliferative disorder.JCI Insight. 2020 Mar 26;5(6):e131644. doi: 10.1172/jci.insight.131644. JCI Insight. 2020. PMID: 32213705 Free PMC article.
-
Genome-Wide Analysis of Epstein-Barr Virus Isolated from Extranodal NK/T-Cell Lymphoma, Nasal Type.Oncologist. 2019 Sep;24(9):e905-e913. doi: 10.1634/theoncologist.2017-0588. Epub 2019 Apr 2. Oncologist. 2019. PMID: 30940744 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

