Legionella pneumophila strain associated with the first evidence of person-to-person transmission of Legionnaires' disease: a unique mosaic genetic backbone
- PMID: 27196677
- PMCID: PMC4872527
- DOI: 10.1038/srep26261
Legionella pneumophila strain associated with the first evidence of person-to-person transmission of Legionnaires' disease: a unique mosaic genetic backbone
Abstract
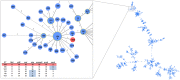
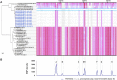
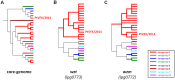
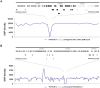
A first strong evidence of person-to-person transmission of Legionnaires' Disease (LD) was recently reported. Here, we characterize the genetic backbone of this case-related Legionella pneumophila strain ("PtVFX/2014"), which also caused a large outbreak of LD. PtVFX/2014 is phylogenetically divergent from the most worldwide studied outbreak-associated L. pneumophila subspecies pneumophila serogroup 1 strains. In fact, this strain is also from serogroup 1, but belongs to the L. pneumophila subspecies fraseri. Its genomic mosaic backbone reveals eight horizontally transferred regions encompassing genes, for instance, involved in lipopolysaccharide biosynthesis or encoding virulence-associated Dot/Icm type IVB secretion system (T4BSS) substrates. PtVFX/2014 also inherited a rare ~65 kb pathogenicity island carrying virulence factors and detoxifying enzymes believed to contribute to the emergence of best-fitted strains in water reservoirs and in human macrophages, as well as a inter-species transferred (from L. oakridgensis) ~37.5 kb genomic island (harboring a lvh/lvr T4ASS cluster) that had never been found intact within L. pneumophila species. PtVFX/2014 encodes another lvh/lvr cluster near to CRISPR-associated genes, which may boost L. pneumophila transition from an environmental bacterium to a human pathogen. Overall, this unique genomic make-up may impact PtVFX/2014 ability to adapt to diverse environments, and, ultimately, to be transmitted and cause human disease.
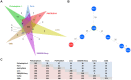
Figures
Similar articles
-
Comparative genome analysis reveals a complex population structure of Legionella pneumophila subspecies.Infect Genet Evol. 2018 Apr;59:172-185. doi: 10.1016/j.meegid.2018.02.008. Epub 2018 Feb 7. Infect Genet Evol. 2018. PMID: 29427765 Free PMC article.
-
Subtyping of the Legionella pneumophila "Ulm" outbreak strain using the CRISPR-Cas system.Int J Med Microbiol. 2015 Dec;305(8):828-37. doi: 10.1016/j.ijmm.2015.08.001. Epub 2015 Aug 7. Int J Med Microbiol. 2015. PMID: 26294350
-
Gene flow in environmental Legionella pneumophila leads to genetic and pathogenic heterogeneity within a Legionnaires' disease outbreak.Genome Biol. 2014;15(11):504. doi: 10.1186/PREACCEPT-1675723368141690. Genome Biol. 2014. PMID: 25370747 Free PMC article.
-
Molecular epidemiology, phylogeny and evolution of Legionella.Infect Genet Evol. 2016 Sep;43:108-22. doi: 10.1016/j.meegid.2016.04.033. Epub 2016 May 13. Infect Genet Evol. 2016. PMID: 27180896 Review.
-
Legionella pneumophila: an aquatic microbe goes astray.FEMS Microbiol Rev. 2002 Jun;26(2):149-62. doi: 10.1111/j.1574-6976.2002.tb00607.x. FEMS Microbiol Rev. 2002. PMID: 12069880 Review.
Cited by
-
Development of heat-shock resistance in Legionella pneumophila modeled by experimental evolution.Appl Environ Microbiol. 2023 Sep 28;89(9):e0066623. doi: 10.1128/aem.00666-23. Epub 2023 Sep 5. Appl Environ Microbiol. 2023. PMID: 37668382 Free PMC article.
-
Recurrence, Microevolution, and Spatiotemporal Dynamics of Legionella pneumophila Sequence Type 1905, Portugal, 2014-2022.Emerg Infect Dis. 2024 May;30(5):1022-1025. doi: 10.3201/eid3005.231383. Emerg Infect Dis. 2024. PMID: 38666647 Free PMC article.
-
Review Global seroprevalence of legionellosis - a systematic review and meta-analysis.Sci Rep. 2020 Apr 30;10(1):7337. doi: 10.1038/s41598-020-63740-y. Sci Rep. 2020. PMID: 32355282 Free PMC article.
-
Complete Genome Sequences of Legionella pneumophila subsp. fraseri Strains Detroit-1 and Dallas 1E.Genome Announc. 2017 Feb 2;5(5):e01525-16. doi: 10.1128/genomeA.01525-16. Genome Announc. 2017. PMID: 28153889 Free PMC article.
-
Atypical Pneumonia: Updates on Legionella, Chlamydophila, and Mycoplasma Pneumonia.Clin Chest Med. 2017 Mar;38(1):45-58. doi: 10.1016/j.ccm.2016.11.011. Epub 2016 Dec 24. Clin Chest Med. 2017. PMID: 28159161 Free PMC article. Review.
References
-
- Dominguez A. et al. Factors influencing the case-fatality rate of Legionnaires’ disease. Int J Tuberc Lung Dis 13, 407–412 (2009). - PubMed
-
- Phin N. et al. Epidemiology and clinical management of Legionnaires’ disease. Lancet Infect Dis 14, 1011–1021 (2014). - PubMed
-
- Yu V. L. et al. Distribution of Legionella species and serogroups isolated by culture in patients with sporadic community-acquired legionellosis: an international collaborative survey. J Infect Dis 186, 127–128 (2002). - PubMed
-
- Centers for Disease Control and Prevention. Legionellosis–United States, 2000–2009. MMWR Morb Mortal Wkly Rep 60, 1083–1086 (2011). - PubMed
-
- European Centre for Disease Prevention and Control. Legionnaires’ disease in Europe, 2012. (ECDC, Stockholm, 2014).
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

