Weight Loss Upregulates the Small GTPase DIRAS3 in Human White Adipose Progenitor Cells, Which Negatively Regulates Adipogenesis and Activates Autophagy via Akt-mTOR Inhibition
- PMID: 27211557
- PMCID: PMC4856797
- DOI: 10.1016/j.ebiom.2016.03.030
Weight Loss Upregulates the Small GTPase DIRAS3 in Human White Adipose Progenitor Cells, Which Negatively Regulates Adipogenesis and Activates Autophagy via Akt-mTOR Inhibition
Abstract
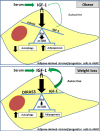
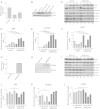
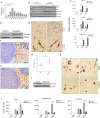
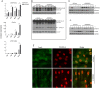
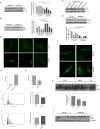
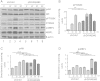
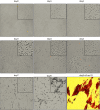
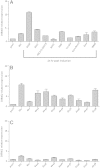
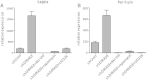
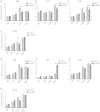
Long-term weight-loss (WL) interventions reduce insulin serum levels, protect from obesity, and postpone age-associated diseases. The impact of long-term WL on adipose-derived stromal/progenitor cells (ASCs) is unknown. We identified DIRAS3 and IGF-1 as long-term WL target genes up-regulated in ASCs in subcutaneous white adipose tissue of formerly obese donors (WLDs). We show that DIRAS3 negatively regulates Akt, mTOR and ERK1/2 signaling in ASCs undergoing adipogenesis and acts as a negative regulator of this pathway and an activator of autophagy. Studying the IGF-1-DIRAS3 interaction in ASCs of WLDs, we demonstrate that IGF-1, although strongly up-regulated in these cells, hardly activates Akt, while ERK1/2 and S6K1 phosphorylation is activated by IGF-1. Overexpression of DIRAS3 in WLD ASCs completely inhibits Akt phosphorylation also in the presence of IGF-1. Phosphorylation of ERK1/2 and S6K1 is lesser reduced under these conditions. In conclusion, our key findings are that DIRAS3 down-regulates Akt-mTOR signaling in ASCs of WLDs. Moreover, DIRAS3 inhibits adipogenesis and activates autophagy in these cells.
Keywords: Adipogenesis; Aging; Akt; Autophagy; Caloric restriction; DIRAS3; ERK1/2; Human adipose-derived stromal/progenitor cells; IGF-1; Insulin; Obesity; Weight loss; mTOR.
Copyright © 2016 The Authors. Published by Elsevier B.V. All rights reserved.
Figures





Similar articles
-
Silencing of the small GTPase DIRAS3 induces cellular senescence in human white adipose stromal/progenitor cells.Aging (Albany NY). 2017 Mar 17;9(3):860-879. doi: 10.18632/aging.101197. Aging (Albany NY). 2017. PMID: 28316325 Free PMC article.
-
The tumour suppressor DiRas3 interacts with C-RAF and downregulates MEK activity to restrict cell migration.Biol Cell. 2013 Feb;105(2):91-107. doi: 10.1111/boc.201200030. Epub 2012 Dec 28. Biol Cell. 2013. PMID: 23157514
-
Bariatric surgery and diet-induced long-term caloric restriction protect subcutaneous adipose-derived stromal/progenitor cells and prolong their life span in formerly obese humans.Exp Gerontol. 2014 Aug;56:106-13. doi: 10.1016/j.exger.2014.03.030. Epub 2014 Apr 18. Exp Gerontol. 2014. PMID: 24747059
-
Protein restriction and cancer.Biochim Biophys Acta Rev Cancer. 2018 Apr;1869(2):256-262. doi: 10.1016/j.bbcan.2018.03.004. Epub 2018 Mar 26. Biochim Biophys Acta Rev Cancer. 2018. PMID: 29596961 Review.
-
p53 regulation of the IGF-1/AKT/mTOR pathways and the endosomal compartment.Cold Spring Harb Perspect Biol. 2010 Feb;2(2):a001057. doi: 10.1101/cshperspect.a001057. Cold Spring Harb Perspect Biol. 2010. PMID: 20182617 Free PMC article. Review.
Cited by
-
Chronic everolimus treatment of high-fat diet mice leads to a reduction in obesity but impaired glucose tolerance.Pharmacol Res Perspect. 2021 Apr;9(2):e00732. doi: 10.1002/prp2.732. Pharmacol Res Perspect. 2021. PMID: 33715287 Free PMC article.
-
An organoid model derived from human adipose stem/progenitor cells to study adipose tissue physiology.Adipocyte. 2022 Dec;11(1):164-174. doi: 10.1080/21623945.2022.2044601. Adipocyte. 2022. PMID: 35297273 Free PMC article.
-
Metformin Improves Stemness of Human Adipose-Derived Stem Cells by Downmodulation of Mechanistic Target of Rapamycin (mTOR) and Extracellular Signal-Regulated Kinase (ERK) Signaling.Biomedicines. 2021 Nov 27;9(12):1782. doi: 10.3390/biomedicines9121782. Biomedicines. 2021. PMID: 34944598 Free PMC article.
-
Autophagy in Stem Cell Biology: A Perspective on Stem Cell Self-Renewal and Differentiation.Stem Cells Int. 2018 Jan 21;2018:9131397. doi: 10.1155/2018/9131397. eCollection 2018. Stem Cells Int. 2018. PMID: 29765428 Free PMC article. Review.
-
An optimized pipeline for high-throughput bulk RNA-Seq deconvolution illustrates the impact of obesity and weight loss on cell composition of human adipose tissue.bioRxiv [Preprint]. 2024 Sep 25:2024.09.23.614489. doi: 10.1101/2024.09.23.614489. bioRxiv. 2024. Update in: Obesity (Silver Spring). 2025 May;33(5):936-948. doi: 10.1002/oby.24264. PMID: 39386599 Free PMC article. Updated. Preprint.
References
-
- Bäck K., Arnqvist H.J. Changes in insulin and IGF-I receptor expression during differentiation of human preadipocytes. Growth Hormon. IGF Res. 2009;19:101–111. - PubMed
-
- Blüher M., Michael M.D., Peroni O.D., Ueki K., Carter N., Kahn B.B., Kahn C.R. Adipose tissue selective insulin receptor knockout protects against obesity and obesity-related glucose intolerance. Dev. Cell. 2002;3:25–38. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

