Characterizing Strain Variation in Engineered E. coli Using a Multi-Omics-Based Workflow
- PMID: 27211860
- PMCID: PMC4882250
- DOI: 10.1016/j.cels.2016.04.004
Characterizing Strain Variation in Engineered E. coli Using a Multi-Omics-Based Workflow
Abstract
Understanding the complex interactions that occur between heterologous and native biochemical pathways represents a major challenge in metabolic engineering and synthetic biology. We present a workflow that integrates metabolomics, proteomics, and genome-scale models of Escherichia coli metabolism to study the effects of introducing a heterologous pathway into a microbial host. This workflow incorporates complementary approaches from computational systems biology, metabolic engineering, and synthetic biology; provides molecular insight into how the host organism microenvironment changes due to pathway engineering; and demonstrates how biological mechanisms underlying strain variation can be exploited as an engineering strategy to increase product yield. As a proof of concept, we present the analysis of eight engineered strains producing three biofuels: isopentenol, limonene, and bisabolene. Application of this workflow identified the roles of candidate genes, pathways, and biochemical reactions in observed experimental phenomena and facilitated the construction of a mutant strain with improved productivity. The contributed workflow is available as an open-source tool in the form of iPython notebooks.
Copyright © 2016 Elsevier Inc. All rights reserved.
Figures


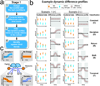




Comment in
-
Omics Meets Metabolic Pathway Engineering.Cell Syst. 2016 Jun 22;2(6):362-3. doi: 10.1016/j.cels.2016.05.005. Epub 2016 May 26. Cell Syst. 2016. PMID: 27237740
References
-
- Almaas E, Kovács B, Vicsek T, Oltvai ZN, Barabási A-L. Global organization of metabolic fluxes in the bacterium Escherichia coli. Nature. 2004;427:839–843. - PubMed
-
- Alonso-Gutierrez J, Chan R, Batth TS, Adams PD, Keasling JD, Petzold CJ, Lee TS. Metabolic engineering of Escherichia coli for limonene and perillyl alcohol production. Metab. Eng. 2013;19:33–41. - PubMed
-
- Alonso-Gutierrez J, Kim E-M, Batth TS, Cho N, Hu Q, Chan LJG, Petzold CJ, Hillson NJ, Adams PD, Keasling JD, et al. Principal component analysis of proteomics (PCAP) as a tool to direct metabolic engineering. Metab. Eng. 2015;28:123–133. - PubMed
-
- Amann E, Ochs B, Abel KJ. Tightly regulated tac promoter vectors useful for the expression of unfused and fused proteins in Escherichia coli. Gene. 1988;69:301–315. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

