Characterization of physiological responses to 22 gene knockouts in Escherichia coli central carbon metabolism
- PMID: 27212692
- PMCID: PMC5845446
- DOI: 10.1016/j.ymben.2016.05.006
Characterization of physiological responses to 22 gene knockouts in Escherichia coli central carbon metabolism
Abstract
Understanding the impact of gene knockouts on cellular physiology, and metabolism in particular, is centrally important to quantitative systems biology and metabolic engineering. Here, we present a comprehensive physiological characterization of wild-type Escherichia coli and 22 knockouts of enzymes in the upper part of central carbon metabolism, including the PTS system, glycolysis, pentose phosphate pathway and Entner-Doudoroff pathway. Our results reveal significant metabolic changes that are affected by specific gene knockouts. Analysis of collective trends and correlations in the data using principal component analysis (PCA) provide new, and sometimes surprising, insights into E. coli physiology. Additionally, by comparing the data-to-model predictions from constraint-based approaches such as FBA, MOMA and RELATCH we demonstrate the important role of less well-understood kinetic and regulatory effects in central carbon metabolism.
Keywords: COBRA modeling; Cell physiology; Escherichia coli; Gene knockout; Metabolism.
Copyright © 2016 International Metabolic Engineering Society. Published by Elsevier Inc. All rights reserved.
Figures





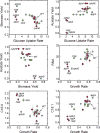


References
-
- Au J, Choi J, Jones SW, Venkataramanan KP, Antoniewicz MR. Parallel labeling experiments validate Clostridium acetobutylicum metabolic network model for (13)C metabolic flux analysis. Metab. Eng. 2014;26C:23–33. - PubMed
-
- Bailey JE. Toward a Science of Metabolic Engineering. Science. 1991;252:1668–1675. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

