Establishment of a Conditionally Immortalized Wilms Tumor Cell Line with a Homozygous WT1 Deletion within a Heterozygous 11p13 Deletion and UPD Limited to 11p15
- PMID: 27213811
- PMCID: PMC4876997
- DOI: 10.1371/journal.pone.0155561
Establishment of a Conditionally Immortalized Wilms Tumor Cell Line with a Homozygous WT1 Deletion within a Heterozygous 11p13 Deletion and UPD Limited to 11p15
Abstract
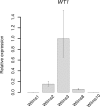
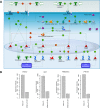
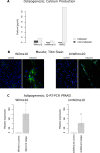
We describe a stromal predominant Wilms tumor with focal anaplasia and a complex, tumor specific chromosome 11 aberration: a homozygous deletion of the entire WT1 gene within a heterozygous 11p13 deletion and an additional region of uniparental disomy (UPD) limited to 11p15.5-p15.2 including the IGF2 gene. The tumor carried a heterozygous p.T41A mutation in CTNNB1. Cells established from the tumor carried the same chromosome 11 aberration, but a different, homozygous p.S45Δ CTNNB1 mutation. Uniparental disomy (UPD) 3p21.3pter lead to the homozygous CTNNB1 mutation. The tumor cell line was immortalized using the catalytic subunit of human telomerase (hTERT) in conjunction with a novel thermolabile mutant (U19dl89-97tsA58) of SV40 large T antigen (LT). This cell line is cytogenetically stable and can be grown indefinitely representing a valuable tool to study the effect of a complete lack of WT1 in tumor cells. The origin/fate of Wilms tumors with WT1 mutations is currently poorly defined. Here we studied the expression of several genes expressed in early kidney development, e.g. FOXD1, PAX3, SIX1, OSR1, OSR2 and MEIS1 and show that these are expressed at similar levels in the parental and the immortalized Wilms10 cells. In addition the limited potential for muscle/ osteogenic/ adipogenic differentiation similar to all other WT1 mutant cell lines is also observed in the Wilms10 tumor cell line and this is retained in the immortalized cells. In summary these Wilms10 cells are a valuable model system for functional studies of WT1 mutant cells.
Conflict of interest statement
Figures






Similar articles
-
Stratification of Wilms tumor by genetic and epigenetic analysis.Oncotarget. 2012 Mar;3(3):327-35. doi: 10.18632/oncotarget.468. Oncotarget. 2012. PMID: 22470196 Free PMC article.
-
Infrequent mutation of the WT1 gene in 77 Wilms' Tumors.Hum Mutat. 1994;3(3):212-22. doi: 10.1002/humu.1380030307. Hum Mutat. 1994. PMID: 8019557
-
Deletion of WT1 and WIT1 genes and loss of heterozygosity on chromosome 11p in Wilms tumors in Japan.Jpn J Cancer Res. 1993 Jun;84(6):616-24. doi: 10.1111/j.1349-7006.1993.tb02021.x. Jpn J Cancer Res. 1993. PMID: 8393432 Free PMC article.
-
The role of Wilms' tumor genes.J Med Invest. 1999 Aug;46(3-4):130-40. J Med Invest. 1999. PMID: 10687307 Review.
-
WT1: a novel tumor suppressor gene inactivated in Wilms' tumor.New Biol. 1992 Feb;4(2):97-106. New Biol. 1992. PMID: 1313285 Review.
Cited by
-
Exploiting embryonic niche conditions to grow Wilms tumor blastema in culture.Front Oncol. 2023 Mar 16;13:1091274. doi: 10.3389/fonc.2023.1091274. eCollection 2023. Front Oncol. 2023. PMID: 37007076 Free PMC article.
-
Comprehensive Biology and Genetics Compendium of Wilms Tumor Cell Lines with Different WT1 Mutations.Cancers (Basel). 2020 Dec 28;13(1):60. doi: 10.3390/cancers13010060. Cancers (Basel). 2020. PMID: 33379206 Free PMC article.
-
Gene expression studies of WT1 mutant Wilms tumor cell lines in the frame work of published kidney development data reveals their early kidney stem cell origin.PLoS One. 2023 Jan 23;18(1):e0270380. doi: 10.1371/journal.pone.0270380. eCollection 2023. PLoS One. 2023. PMID: 36689432 Free PMC article.
-
Chromosomal Heterogeneity of the G-401 Rhabdoid Tumor Cell Line: Unusual Partial 7p Trisomy.Front Med (Lausanne). 2019 Aug 30;6:187. doi: 10.3389/fmed.2019.00187. eCollection 2019. Front Med (Lausanne). 2019. PMID: 31544104 Free PMC article.
-
Targeting TRIP13 in favorable histology Wilms tumor with nuclear export inhibitors synergizes with doxorubicin.Commun Biol. 2024 Apr 8;7(1):426. doi: 10.1038/s42003-024-06140-6. Commun Biol. 2024. PMID: 38589567 Free PMC article.
References
-
- Maiti S, Alam R, Amos CI, Huff V. (2000) Frequent association of β-catenin and WT1 mutations in Wilms tumors. Cancer Res 60: 6288–6292. - PubMed
-
- Schumacher V, Schuhen S, Sonner S, Weirich A, Leuschner I, Harms D, et al. (2003) Two molecular subgroups of Wilms' tumors with or without WT1 mutations. Clin Cancer Res 9: 2005–2014. - PubMed
-
- Little SE, Hanks SP, Kind-Underwood L, Jones C, Rapley EA, Rahman N, et al. (2004) Frequency and heritability of WT1 mutations in nonsyndromic Wilms’ tumor patients: A UK Children’s Cancer Study Group Study. J Clin Oncol 22: 4140–4146. - PubMed
-
- Royer-Pokora B, Weirich A, Schumacher V, Uschkereit C, Beier M, Leuschner I, et al. (2008) Clinical relevance of mutations in the Wilms tumor suppressor 1 gene WT1 and the cadherin-associated protein beta1 gene CTNNB1 for patients with Wilms tumors: results of long-term surveillance of 71 patients from International Society of Pediatric Oncology Study 9/Society for Pediatric Oncology Cancer 5: 1080–1089. - PubMed
Publication types
MeSH terms
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

