SWIFT-Review: a text-mining workbench for systematic review
- PMID: 27216467
- PMCID: PMC4877757
- DOI: 10.1186/s13643-016-0263-z
SWIFT-Review: a text-mining workbench for systematic review
Abstract
Background: There is growing interest in using machine learning approaches to priority rank studies and reduce human burden in screening literature when conducting systematic reviews. In addition, identifying addressable questions during the problem formulation phase of systematic review can be challenging, especially for topics having a large literature base. Here, we assess the performance of the SWIFT-Review priority ranking algorithm for identifying studies relevant to a given research question. We also explore the use of SWIFT-Review during problem formulation to identify, categorize, and visualize research areas that are data rich/data poor within a large literature corpus.
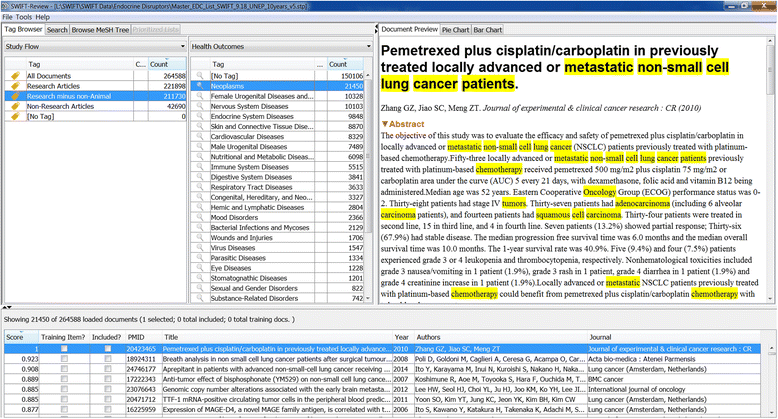
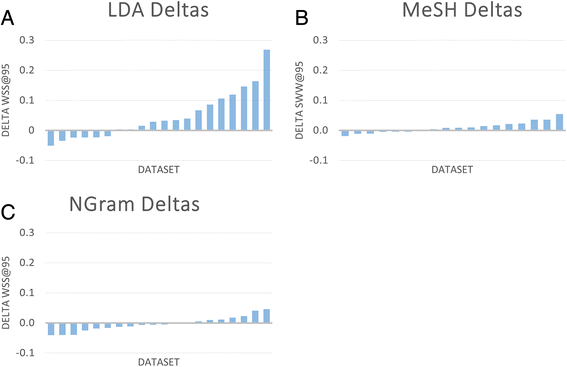
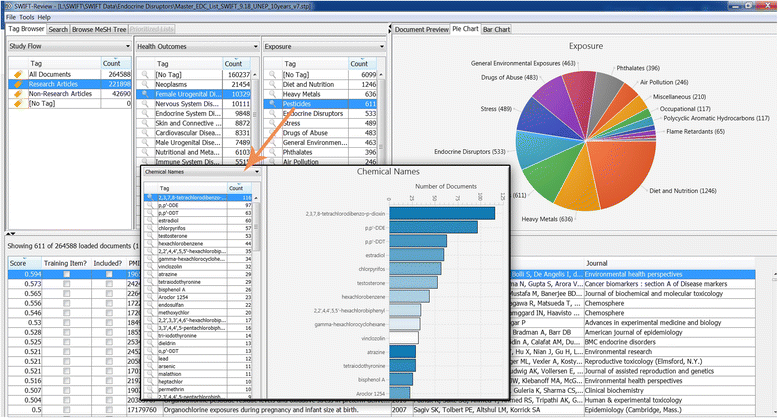
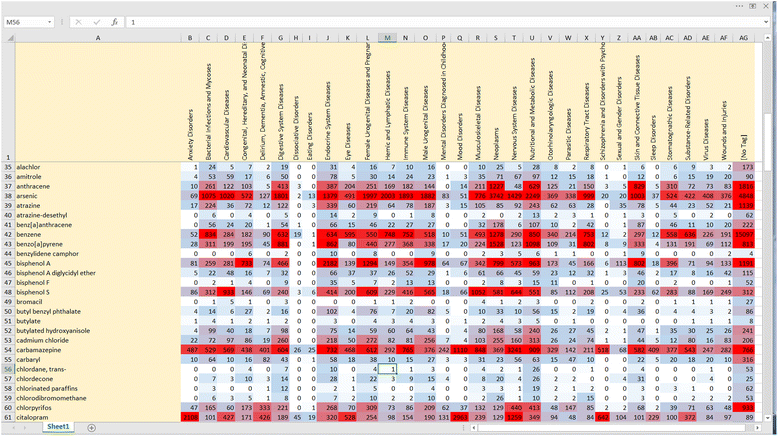
Methods: Twenty case studies, including 15 public data sets, representing a range of complexity and size, were used to assess the priority ranking performance of SWIFT-Review. For each study, seed sets of manually annotated included and excluded titles and abstracts were used for machine training. The remaining references were then ranked for relevance using an algorithm that considers term frequency and latent Dirichlet allocation (LDA) topic modeling. This ranking was evaluated with respect to (1) the number of studies screened in order to identify 95 % of known relevant studies and (2) the "Work Saved over Sampling" (WSS) performance metric. To assess SWIFT-Review for use in problem formulation, PubMed literature search results for 171 chemicals implicated as EDCs were uploaded into SWIFT-Review (264,588 studies) and categorized based on evidence stream and health outcome. Patterns of search results were surveyed and visualized using a variety of interactive graphics.
Results: Compared with the reported performance of other tools using the same datasets, the SWIFT-Review ranking procedure obtained the highest scores on 11 out of 15 of the public datasets. Overall, these results suggest that using machine learning to triage documents for screening has the potential to save, on average, more than 50 % of the screening effort ordinarily required when using un-ordered document lists. In addition, the tagging and annotation capabilities of SWIFT-Review can be useful during the activities of scoping and problem formulation.
Conclusions: Text-mining and machine learning software such as SWIFT-Review can be valuable tools to reduce the human screening burden and assist in problem formulation.
Keywords: Literature prioritization; SWIFT-Review; Scoping reports; Software; Systematic review.
Figures





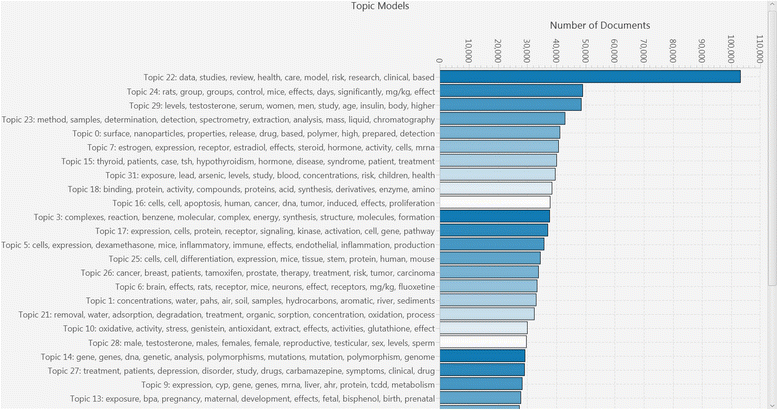
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

