Whole genome sequencing for deciphering the resistome of Chryseobacterium indologenes, an emerging multidrug-resistant bacterium isolated from a cystic fibrosis patient in Marseille, France
- PMID: 27222716
- PMCID: PMC4873609
- DOI: 10.1016/j.nmni.2016.03.006
Whole genome sequencing for deciphering the resistome of Chryseobacterium indologenes, an emerging multidrug-resistant bacterium isolated from a cystic fibrosis patient in Marseille, France
Abstract
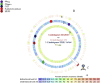
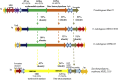
We decipher the resistome of Chryseobacterium indologenes MARS15, an emerging multidrug-resistant clinical strain, using the whole genome sequencing strategy. The bacterium was isolated from the sputum of a hospitalized patient with cystic fibrosis in the Timone Hospital in Marseille, France. Genome sequencing was done with Illumina MiSeq using a paired-end strategy. The in silico analysis was done by RAST, the resistome by the ARG-ANNOT database and detection of polyketide synthase (PKS) by ANTISMAH. The genome size of C. indologenes MARS15 is 4 972 580 bp with 36.4% GC content. This multidrug-resistant bacterium was resistant to all β-lactams, including imipenem, and also to colistin. The resistome of C. indologenes MARS15 includes Ambler class A and B β-lactams encoding bla CIA and bla IND-2 genes and MBL (metallo-β-lactamase) genes, the CAT (chloramphenicol acetyltransferase) gene and the multidrug efflux pump AcrB. Specific features include the presence of an urease operon, an intact prophage and a carotenoid biosynthesis pathway. Interestingly, we report for the first time in C. indologenes a PKS cluster that might be responsible for secondary metabolite biosynthesis, similar to erythromycin. The whole genome sequence analysis provides insight into the resistome and the discovery of new details, such as the PKS cluster.
Keywords: Carotenoid; Chryseobacterium indologenes; microbial genomics; nonribosomal polyketide synthase; polyketide synthase; resistome; urease.
Figures


Similar articles
-
Whole-genome sequence of Chryseobacterium oranimense, a colistin-resistant bacterium isolated from a cystic fibrosis patient in France.Antimicrob Agents Chemother. 2015 Mar;59(3):1696-706. doi: 10.1128/AAC.02417-14. Epub 2015 Jan 12. Antimicrob Agents Chemother. 2015. PMID: 25583710 Free PMC article.
-
Whole genome sequencing of the multidrug-resistant Chryseobacterium indologenes isolated from a patient in Brazil.Front Med (Lausanne). 2022 Jul 28;9:931379. doi: 10.3389/fmed.2022.931379. eCollection 2022. Front Med (Lausanne). 2022. PMID: 35966843 Free PMC article.
-
Whole genome sequencing uncovers a novel IND-16 metallo-β-lactamase from an extensively drug-resistant Chryseobacterium indologenes strain J31.Gut Pathog. 2016 Oct 21;8:47. doi: 10.1186/s13099-016-0130-4. eCollection 2016. Gut Pathog. 2016. PMID: 27785154 Free PMC article.
-
Is Chryseobacterium indologenes a shunt-lover bacterium? A case report and review of the literature.Infez Med. 2013 Dec;21(4):312-6. Infez Med. 2013. PMID: 24335463 Review.
-
Six cases during 2012-2015 and literature review of Chryseobacterium indologenes infections in pediatric patients.Can J Microbiol. 2016 Oct;62(10):812-819. doi: 10.1139/cjm-2015-0800. Epub 2016 May 17. Can J Microbiol. 2016. PMID: 27397741 Review.
Cited by
-
Rarely Encountered Gram-Negative Rods and Lung Transplant Recipients: A Narrative Review.Microorganisms. 2023 May 31;11(6):1468. doi: 10.3390/microorganisms11061468. Microorganisms. 2023. PMID: 37374970 Free PMC article. Review.
-
Phylogenetic insights into the diversity of Chryseobacterium species.Access Microbiol. 2019 May 3;1(3):e000019. doi: 10.1099/acmi.0.000019. eCollection 2019. Access Microbiol. 2019. PMID: 32974515 Free PMC article.
-
Detection of 5 Kinds of Genes Related to Plasmid-Mediated Quinolone Resistance in Four Species of Nonfermenting Bacteria with 2 Drug Resistant Phenotypes.Can J Infect Dis Med Microbiol. 2020 Apr 9;2020:3948719. doi: 10.1155/2020/3948719. eCollection 2020. Can J Infect Dis Med Microbiol. 2020. PMID: 32351636 Free PMC article.
-
In vitro assays reveal inherently insecticide-tolerant termite symbionts.Front Physiol. 2023 Jul 12;14:1134936. doi: 10.3389/fphys.2023.1134936. eCollection 2023. Front Physiol. 2023. PMID: 37501931 Free PMC article.
-
Challenges in the identification of Chryseobacterium indologenes and Elizabethkingia meningoseptica in cases of nosocomial infections and patients with cystic fibrosis.New Microbes New Infect. 2017 Sep 13;20:27-33. doi: 10.1016/j.nmni.2017.09.002. eCollection 2017 Nov. New Microbes New Infect. 2017. PMID: 29062487 Free PMC article.
References
-
- Vandamme P., Bernardet J.F., Segers P., Kersters K., Holmes B. New perspectives in the classification of the flavobacteria: description of Chryseobacterium gen. nov., Bergeyella gen. nov., and Empedobacter nom. rev. Int J Syst Bacteriol. 1994;44:827–831.
-
- Deng L., Li M.F., Li Y.H., Yang J.L., Zhou X. Chryseobacterium indologenes in four patients with leukemia. Transpl Infect Dis. 2015;17:583–587. - PubMed
-
- Chen X.Y., Zhao R., Chen Z.L., Liu L., Li X.D., Li Y.H. Chryseobacterium polytrichastri sp. nov., isolated from a moss (Polytrichastrum formosum), and emended description of the genus Chryseobacterium. Antonie Van Leeuwenhoek. 2015;107:403–410. - PubMed
-
- Chen F.L., Wang G.C., Teng S.O., Ou T.Y., Yu F.L., Lee W.S. Clinical and epidemiological features of Chryseobacterium indologenes infections: analysis of 215 cases. J Microbiol Immunol Infect. 2013;46:425–432. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
