CRISPR-Mediated Slamf1Δ/Δ Slamf5Δ/Δ Slamf6Δ/Δ Triple Gene Disruption Reveals NKT Cell Defects but Not T Follicular Helper Cell Defects
- PMID: 27223891
- PMCID: PMC4880187
- DOI: 10.1371/journal.pone.0156074
CRISPR-Mediated Slamf1Δ/Δ Slamf5Δ/Δ Slamf6Δ/Δ Triple Gene Disruption Reveals NKT Cell Defects but Not T Follicular Helper Cell Defects
Abstract
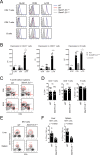
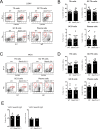
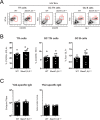
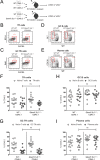
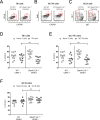
SAP (SH2D1A) is required intrinsically in CD4 T cells to generate germinal center responses and long-term humoral immunity. SAP binds to SLAM family receptors, including SLAM, CD84, and Ly108 to enhance cytokine secretion and sustained T cell:B cell adhesion, which both improve T follicular helper (Tfh) cell aid to germinal center (GC) B cells. To understand the overlapping roles of multiple SLAM family receptors in germinal center responses, Slamf1Δ/Δ Slamf5Δ/Δ Slamf6Δ/Δ triple gene disruption (Slamf1,5,6Δ/Δ) mice were generated using CRISPR-Cas9 gene editing to eliminate expression of SLAM (CD150), CD84, and Ly108, respectively. Gene targeting was highly efficient, with 6 of 6 alleles disrupted in 14 of 23 pups and the majority of alleles disrupted in the remaining pups. NKT cell differentiation in Slamf1,5,6Δ/Δ mice was defective, but not completely absent. The remaining NKT cells exhibited substantially increased 2B4 (SLAMF4) expression. Surprisingly, there were no overt defects in germinal center responses to acute viral infections or protein immunizations in Slamf1,5,6Δ/Δ mice, unlike Sh2d1a-/- mice. Similarly, in the context of a competitive environment, SLAM family receptor expressing GC Tfh cell, GC B cell, and plasma cell responses exhibited no advantages over Slamf1,5,6Δ/Δ cells.
Conflict of interest statement
Figures


Similar articles
-
CRISPR-Mediated Triple Knockout of SLAMF1, SLAMF5 and SLAMF6 Supports Positive Signaling Roles in NKT Cell Development.PLoS One. 2016 Jun 3;11(6):e0156072. doi: 10.1371/journal.pone.0156072. eCollection 2016. PLoS One. 2016. PMID: 27258160 Free PMC article.
-
SAP regulation of follicular helper CD4 T cell development and humoral immunity is independent of SLAM and Fyn kinase.J Immunol. 2007 Jan 15;178(2):817-28. doi: 10.4049/jimmunol.178.2.817. J Immunol. 2007. PMID: 17202343
-
Germinal center T follicular helper cell IL-4 production is dependent on signaling lymphocytic activation molecule receptor (CD150).J Immunol. 2010 Jul 1;185(1):190-202. doi: 10.4049/jimmunol.0903505. Epub 2010 Jun 4. J Immunol. 2010. PMID: 20525889 Free PMC article.
-
The Role of Adaptor Proteins in the Biology of Natural Killer T (NKT) Cells.Front Immunol. 2019 Jun 25;10:1449. doi: 10.3389/fimmu.2019.01449. eCollection 2019. Front Immunol. 2019. PMID: 31293596 Free PMC article. Review.
-
Modulation of SAP dependent T:B cell interactions as a strategy to improve vaccination.Curr Opin Virol. 2013 Jun;3(3):363-70. doi: 10.1016/j.coviro.2013.05.015. Epub 2013 Jun 3. Curr Opin Virol. 2013. PMID: 23743125 Free PMC article. Review.
Cited by
-
Regulatory Effect of Sishen Pill on Tfh Cells in Mice With Experimental Colitis.Front Physiol. 2020 Jun 5;11:589. doi: 10.3389/fphys.2020.00589. eCollection 2020. Front Physiol. 2020. PMID: 32581849 Free PMC article.
-
SLAM-SAP-Fyn: Old Players with New Roles in iNKT Cell Development and Function.Int J Mol Sci. 2019 Sep 27;20(19):4797. doi: 10.3390/ijms20194797. Int J Mol Sci. 2019. PMID: 31569599 Free PMC article. Review.
-
Critical Role for SLAM/SAP Signaling in the Thymic Developmental Programming of IL-17- and IFN-γ-Producing γδ T Cells.J Immunol. 2020 Mar 15;204(6):1521-1534. doi: 10.4049/jimmunol.1901082. Epub 2020 Feb 5. J Immunol. 2020. PMID: 32024701 Free PMC article.
-
SLAM family receptors control pro-survival effectors in germinal center B cells to promote humoral immunity.J Exp Med. 2021 Mar 1;218(3):e20200756. doi: 10.1084/jem.20200756. J Exp Med. 2021. PMID: 33237304 Free PMC article.
-
The Protein Phosphatase Shp1 Regulates Invariant NKT Cell Effector Differentiation Independently of TCR and Slam Signaling.J Immunol. 2019 Apr 15;202(8):2276-2286. doi: 10.4049/jimmunol.1800844. Epub 2019 Feb 22. J Immunol. 2019. PMID: 30796181 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

