Coordinated control of Notch/Delta signalling and cell cycle progression drives lateral inhibition-mediated tissue patterning
- PMID: 27226324
- PMCID: PMC4958321
- DOI: 10.1242/dev.134213
Coordinated control of Notch/Delta signalling and cell cycle progression drives lateral inhibition-mediated tissue patterning
Abstract
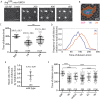
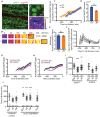
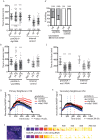
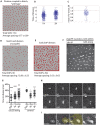
Coordinating cell differentiation with cell growth and division is crucial for the successful development, homeostasis and regeneration of multicellular tissues. Here, we use bristle patterning in the fly notum as a model system to explore the regulatory and functional coupling of cell cycle progression and cell fate decision-making. The pattern of bristles and intervening epithelial cells (ECs) becomes established through Notch-mediated lateral inhibition during G2 phase of the cell cycle, as neighbouring cells physically interact with each other via lateral contacts and/or basal protrusions. Since Notch signalling controls cell division timing downstream of Cdc25, ECs in lateral contact with a Delta-expressing cell experience higher levels of Notch signalling and divide first, followed by more distant neighbours, and lastly Delta-expressing cells. Conversely, mitotic entry and cell division makes ECs refractory to lateral inhibition signalling, fixing their fate. Using a combination of experiments and computational modelling, we show that this reciprocal relationship between Notch signalling and cell cycle progression acts like a developmental clock, providing a delimited window of time during which cells decide their fate, ensuring efficient and orderly bristle patterning.
Keywords: Cell cycle; G2 phase; Lateral inhibition; Notch signalling; Patterning.
© 2016. Published by The Company of Biologists Ltd.
Conflict of interest statement
The authors declare no competing or financial interests.
Figures
Similar articles
-
Par3 cooperates with Sanpodo for the assembly of Notch clusters following asymmetric division of Drosophila sensory organ precursor cells.Elife. 2021 Oct 1;10:e66659. doi: 10.7554/eLife.66659. Elife. 2021. PMID: 34596529 Free PMC article.
-
Self-organized Notch dynamics generate stereotyped sensory organ patterns in Drosophila.Science. 2017 May 5;356(6337):eaai7407. doi: 10.1126/science.aai7407. Epub 2017 Apr 6. Science. 2017. PMID: 28386027
-
Mathematical study of the role of Delta/Notch lateral inhibition during primary branching of Drosophila trachea development.Biophys J. 2012 Dec 19;103(12):2549-59. doi: 10.1016/j.bpj.2012.11.005. Epub 2012 Dec 18. Biophys J. 2012. PMID: 23260057 Free PMC article.
-
Robust selection of sensory organ precursors by the Notch-Delta pathway.Curr Opin Cell Biol. 2011 Dec;23(6):663-7. doi: 10.1016/j.ceb.2011.09.005. Epub 2011 Sep 29. Curr Opin Cell Biol. 2011. PMID: 21963301 Review.
-
Spatio-Temporal Regulation of Notch Activation in Asymmetrically Dividing Sensory Organ Precursor Cells in Drosophila melanogaster Epithelium.Cells. 2024 Jun 30;13(13):1133. doi: 10.3390/cells13131133. Cells. 2024. PMID: 38994985 Free PMC article. Review.
Cited by
-
Immunotherapy and the Combination with Targeted Therapies for Advanced Hepatocellular Carcinoma.Cancers (Basel). 2023 Jan 20;15(3):654. doi: 10.3390/cancers15030654. Cancers (Basel). 2023. PMID: 36765612 Free PMC article. Review.
-
Zebrafish Klf4 maintains the ionocyte progenitor population by regulating epidermal stem cell proliferation and lateral inhibition.PLoS Genet. 2019 Apr 1;15(4):e1008058. doi: 10.1371/journal.pgen.1008058. eCollection 2019 Apr. PLoS Genet. 2019. PMID: 30933982 Free PMC article.
-
A CDK1 phosphorylation site on Drosophila PAR-3 regulates neuroblast polarisation and sensory organ formation.Elife. 2024 Jun 13;13:e97902. doi: 10.7554/eLife.97902. Elife. 2024. PMID: 38869055 Free PMC article.
-
Notch directs telencephalic development and controls neocortical neuron fate determination by regulating microRNA levels.Development. 2023 Jun 1;150(11):dev201408. doi: 10.1242/dev.201408. Epub 2023 Jun 5. Development. 2023. PMID: 37272771 Free PMC article.
-
Cell-Cell Contact Area Affects Notch Signaling and Notch-Dependent Patterning.Dev Cell. 2017 Mar 13;40(5):505-511.e6. doi: 10.1016/j.devcel.2017.02.009. Dev Cell. 2017. PMID: 28292428 Free PMC article.
References
MeSH terms
Substances
Grants and funding
- BB/K009001/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- HHMI/Howard Hughes Medical Institute/United States
- P40 OD018537/OD/NIH HHS/United States
- BB/J008532/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- CRUK_/Cancer Research UK/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

