Generating long-lived CD8(+) T-cell memory: Insights from epigenetic programs
- PMID: 27230488
- PMCID: PMC5035674
- DOI: 10.1002/eji.201545550
Generating long-lived CD8(+) T-cell memory: Insights from epigenetic programs
Abstract
T-cell-based immunological memory has the potential to provide the host with life-long protection against pathogen reexposure and thus offers tremendous promise for the design of vaccines targeting chronic infections or cancer. In order to exploit this potential in the design of new vaccines, it is necessary to understand how and when memory T cells acquire their poised effector potential, and moreover, how they maintain these properties during homeostatic proliferation. To gain insight into the persistent nature of memory T-cell functions, investigators have turned their attention to epigenetic mechanisms. Recent efforts have revealed that many of the properties acquired among memory T cells are coupled to stable changes in DNA methylation and histone modifications. Furthermore, it has recently been reported that the delineating features among memory T cells subsets are also linked to distinct epigenetic events, such as permissive and repressive histone modifications and DNA methylation programs, providing exciting new hypotheses regarding their cellular ancestry. Here, we review recent studies focused on epigenetic programs acquired during effector and memory T-cell differentiation and discuss how these data may shed new light on the developmental path for generating long-lived CD8(+) T-cell memory.
Keywords: CD8+ T cell; DNA methylation; Epigenetics; Histone modification; Memory; T-cell exhaustion.
© 2016 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Figures

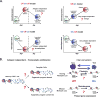
 arrow is reflective of the self-renewing capacity of the memory subsets.
arrow is reflective of the self-renewing capacity of the memory subsets.
Similar articles
-
Epigenetic remodeling of the IL-2 and IFN-gamma loci in memory CD8 T cells is influenced by CD4 T cells.J Immunol. 2006 Jul 15;177(2):1062-9. doi: 10.4049/jimmunol.177.2.1062. J Immunol. 2006. PMID: 16818762
-
Human memory CD8 T cell effector potential is epigenetically preserved during in vivo homeostasis.J Exp Med. 2017 Jun 5;214(6):1593-1606. doi: 10.1084/jem.20161760. Epub 2017 May 10. J Exp Med. 2017. PMID: 28490440 Free PMC article.
-
Remembering to remember: T cell memory maintenance and plasticity.Curr Opin Immunol. 2019 Jun;58:89-97. doi: 10.1016/j.coi.2019.04.009. Epub 2019 Jun 3. Curr Opin Immunol. 2019. PMID: 31170601 Free PMC article. Review.
-
Early transcriptional and epigenetic regulation of CD8+ T cell differentiation revealed by single-cell RNA sequencing.Nat Immunol. 2017 Apr;18(4):422-432. doi: 10.1038/ni.3688. Epub 2017 Feb 20. Nat Immunol. 2017. PMID: 28218746 Free PMC article.
-
The interface between transcriptional and epigenetic control of effector and memory CD8⁺ T-cell differentiation.Immunol Rev. 2014 Sep;261(1):157-68. doi: 10.1111/imr.12205. Immunol Rev. 2014. PMID: 25123283 Free PMC article. Review.
Cited by
-
Exploring Memory Function Beyond Immune Cells: ANGPTL4-Mediated Memory Functions in Tissue Resident Stem Cells.Adv Sci (Weinh). 2024 Jul;11(28):e2307545. doi: 10.1002/advs.202307545. Epub 2024 Apr 26. Adv Sci (Weinh). 2024. PMID: 38666393 Free PMC article.
-
Genetic mechanisms underlying tumor microenvironment composition and function in diffuse large B-cell lymphoma.Blood. 2024 Mar 21;143(12):1101-1111. doi: 10.1182/blood.2023021002. Blood. 2024. PMID: 38211334 Free PMC article. Review.
-
Potential functions of atypical memory B cells in Plasmodium-exposed individuals.Int J Parasitol. 2020 Nov;50(13):1033-1042. doi: 10.1016/j.ijpara.2020.08.003. Epub 2020 Sep 26. Int J Parasitol. 2020. PMID: 32987039 Free PMC article. Review.
-
Identification of memory mechanism in tissue-resident stem cells via ANGPTL4 beyond immune cells upon viral antigen exposure.Mol Ther. 2024 Sep 4;32(9):3042-3058. doi: 10.1016/j.ymthe.2024.04.006. Epub 2024 Apr 6. Mol Ther. 2024. PMID: 38582960
-
Human γδ T cells in diverse tissues exhibit site-specific maturation dynamics across the life span.Sci Immunol. 2024 Jun 7;9(96):eadn3954. doi: 10.1126/sciimmunol.adn3954. Epub 2024 Jun 7. Sci Immunol. 2024. PMID: 38848342 Free PMC article.
References
-
- Alanio C, Lemaitre F, Law HK, Hasan M, Albert ML. Enumeration of human antigen-specific naive CD8+ T cells reveals conserved precursor frequencies. Blood. 2010;115:3718–3725. - PubMed
-
- Coulie PG, Karanikas V, Lurquin C, Colau D, Connerotte T, Hanagiri T, Van Pel A, Lucas S, Godelaine D, Lonchay C, Marchand M, Van Baren N, Boon T. Cytolytic T-cell responses of cancer patients vaccinated with a MAGE antigen. Immunol Rev. 2002;188:33–42. - PubMed
-
- Boon T, Coulie PG, Van den Eynde BJ, van der Bruggen P. Human T cell responses against melanoma. Annu Rev Immunol. 2006;24:175–208. - PubMed
-
- Letsch A, Keilholz U, Schadendorf D, Nagorsen D, Schmittel A, Thiel E, Scheibenbogen C. High frequencies of circulating melanoma-reactive CD8+ T cells in patients with advanced melanoma. Int J Cancer. 2000;87:659–664. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

