Molecular mechanisms regulating formation, trafficking and processing of annular gap junctions
- PMID: 27230503
- PMCID: PMC4896261
- DOI: 10.1186/s12860-016-0087-7
Molecular mechanisms regulating formation, trafficking and processing of annular gap junctions
Abstract
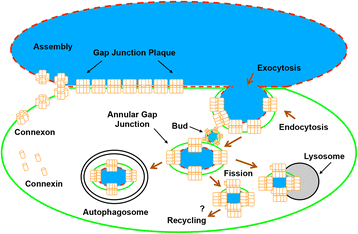
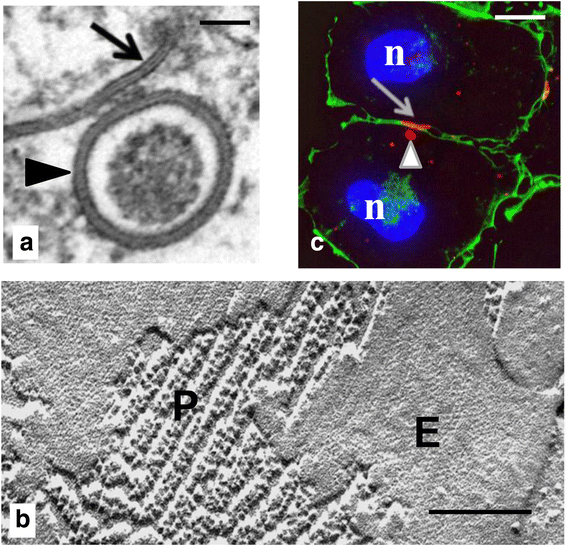
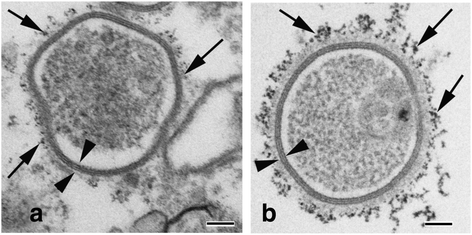
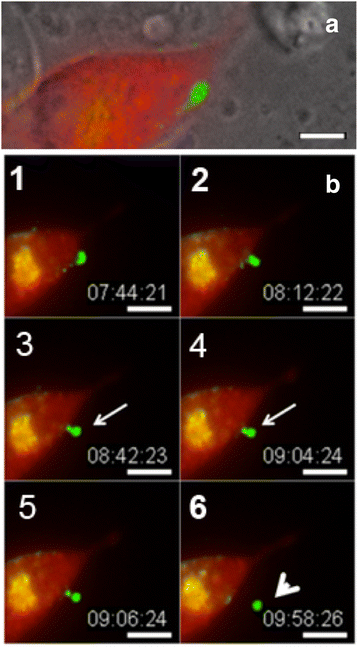
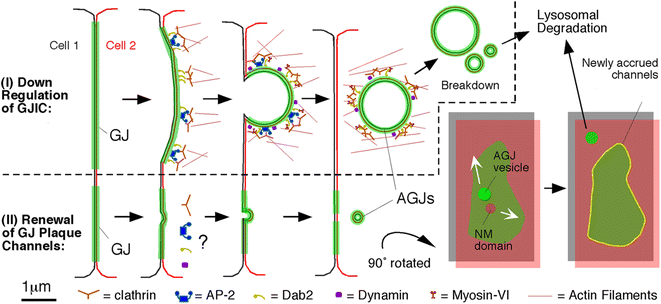
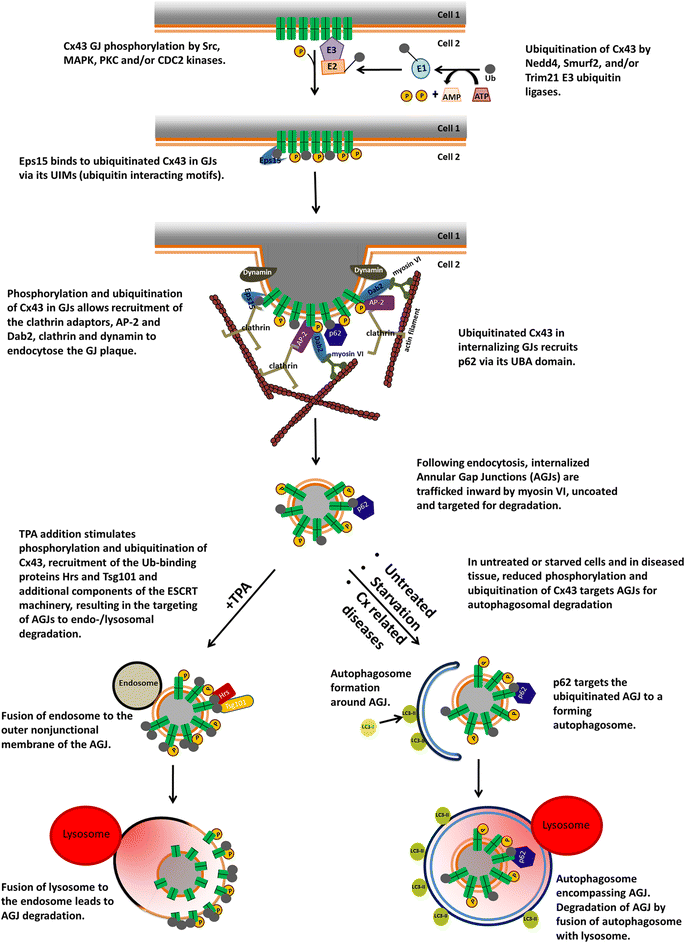
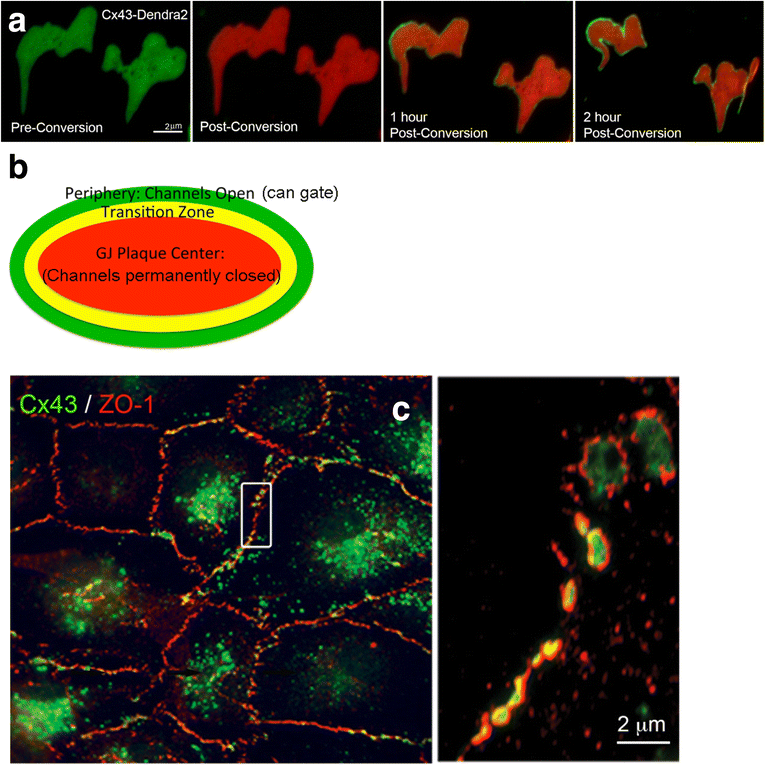
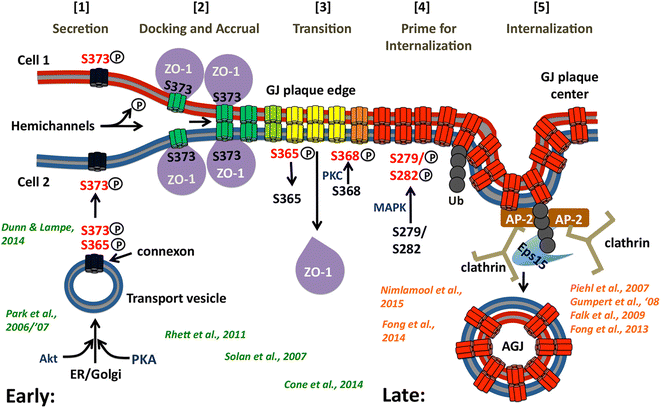
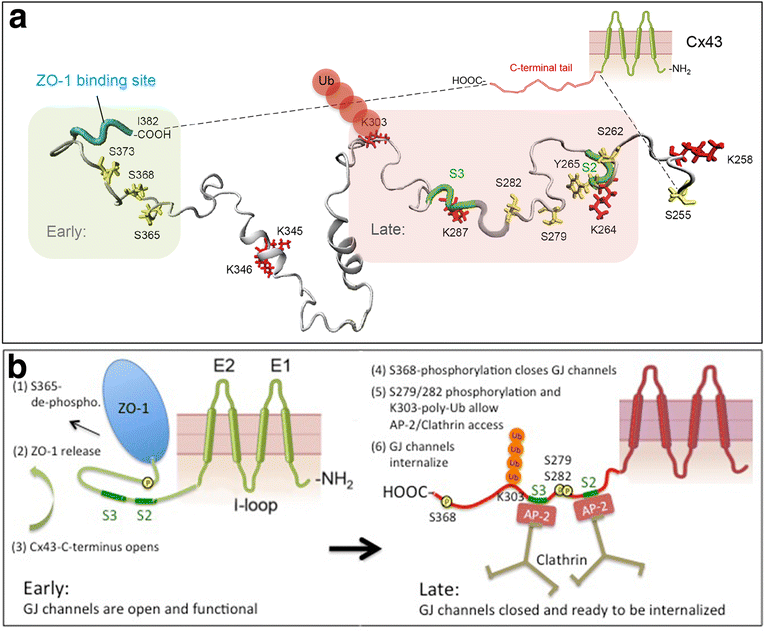
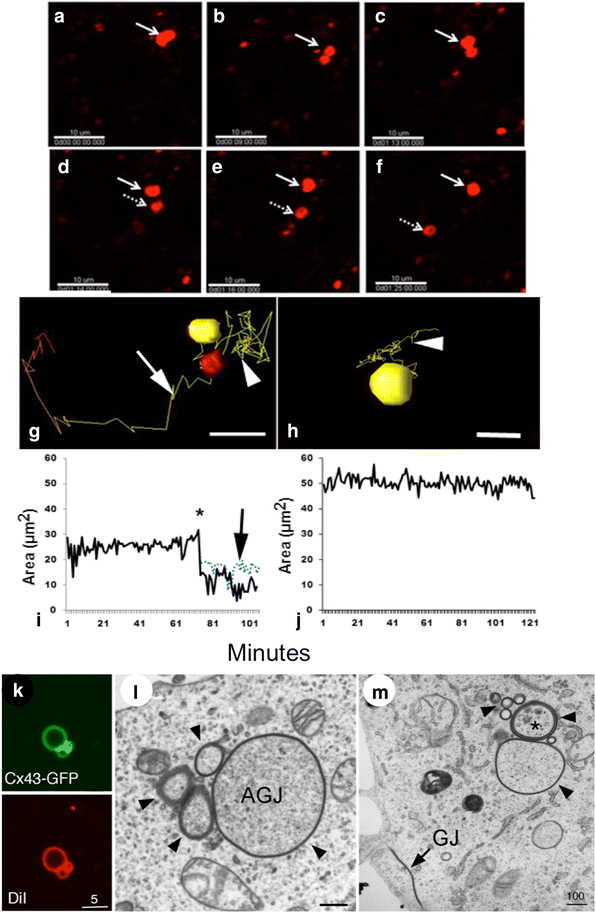
Internalization of gap junction plaques results in the formation of annular gap junction vesicles. The factors that regulate the coordinated internalization of the gap junction plaques to form annular gap junction vesicles, and the subsequent events involved in annular gap junction processing have only relatively recently been investigated in detail. However it is becoming clear that while annular gap junction vesicles have been demonstrated to be degraded by autophagosomal and endo-lysosomal pathways, they undergo a number of additional processing events. Here, we characterize the morphology of the annular gap junction vesicle and review the current knowledge of the processes involved in their formation, fission, fusion, and degradation. In addition, we address the possibility for connexin protein recycling back to the plasma membrane to contribute to gap junction formation and intercellular communication. Information on gap junction plaque removal from the plasma membrane and the subsequent processing of annular gap junction vesicles is critical to our understanding of cell-cell communication as it relates to events regulating development, cell homeostasis, unstable proliferation of cancer cells, wound healing, changes in the ischemic heart, and many other physiological and pathological cellular phenomena.
Keywords: Annular; Clathrin; Connexin; Degradation; Dynamin; Endocytosis; Fission; Gap Junction; Lysosomes; Phosphorylation; Ubiquitination; ZO-1.
Figures


Similar articles
-
Visualizing the effect of dynamin inhibition on annular gap vesicle formation and fission.J Cell Sci. 2013 Jun 15;126(Pt 12):2607-16. doi: 10.1242/jcs.116269. Epub 2013 Apr 16. J Cell Sci. 2013. PMID: 23591819 Free PMC article.
-
Visualization of Annular Gap Junction Vesicle Processing: The Interplay Between Annular Gap Junctions and Mitochondria.Int J Mol Sci. 2018 Dec 22;20(1):44. doi: 10.3390/ijms20010044. Int J Mol Sci. 2018. PMID: 30583492 Free PMC article.
-
Clathrin and Cx43 gap junction plaque endoexocytosis.Biochem Biophys Res Commun. 2008 Oct 3;374(4):679-82. doi: 10.1016/j.bbrc.2008.07.108. Epub 2008 Jul 31. Biochem Biophys Res Commun. 2008. PMID: 18675253
-
Regulation of gap junction intercellular communication by the ubiquitin system.Cell Signal. 2010 Sep;22(9):1267-73. doi: 10.1016/j.cellsig.2010.03.005. Epub 2010 Mar 4. Cell Signal. 2010. PMID: 20206687 Review.
-
Degradation of endocytosed gap junctions by autophagosomal and endo-/lysosomal pathways: a perspective.J Membr Biol. 2012 Aug;245(8):465-76. doi: 10.1007/s00232-012-9464-0. Epub 2012 Jul 24. J Membr Biol. 2012. PMID: 22825714 Free PMC article. Review.
Cited by
-
Regulation of gap junction intercellular communication by connexin ubiquitination: physiological and pathophysiological implications.Cell Mol Life Sci. 2020 Feb;77(4):573-591. doi: 10.1007/s00018-019-03285-0. Epub 2019 Sep 9. Cell Mol Life Sci. 2020. PMID: 31501970 Free PMC article. Review.
-
Multiscale Simulations of Biological Membranes: The Challenge To Understand Biological Phenomena in a Living Substance.Chem Rev. 2019 May 8;119(9):5607-5774. doi: 10.1021/acs.chemrev.8b00538. Epub 2019 Mar 12. Chem Rev. 2019. PMID: 30859819 Free PMC article.
-
The electrical synapse: Molecular complexities at the gap and beyond.Dev Neurobiol. 2017 May;77(5):562-574. doi: 10.1002/dneu.22484. Epub 2017 Mar 8. Dev Neurobiol. 2017. PMID: 28170151 Free PMC article. Review.
-
Suppression of connexin 43 phosphorylation promotes astrocyte survival and vascular regeneration in proliferative retinopathy.Proc Natl Acad Sci U S A. 2018 Jun 26;115(26):E5934-E5943. doi: 10.1073/pnas.1803907115. Epub 2018 Jun 11. Proc Natl Acad Sci U S A. 2018. PMID: 29891713 Free PMC article.
-
Adherens, tight, and gap junctions in ependymal cells: A systematic review of their contribution to CSF-brain barrier.Front Neurol. 2023 Mar 24;14:1092205. doi: 10.3389/fneur.2023.1092205. eCollection 2023. Front Neurol. 2023. PMID: 37034077 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

