Discovery of a Novel Alginate Lyase from Nitratiruptor sp. SB155-2 Thriving at Deep-sea Hydrothermal Vents and Identification of the Residues Responsible for Its Heat Stability
- PMID: 27231344
- PMCID: PMC4957041
- DOI: 10.1074/jbc.M115.713230
Discovery of a Novel Alginate Lyase from Nitratiruptor sp. SB155-2 Thriving at Deep-sea Hydrothermal Vents and Identification of the Residues Responsible for Its Heat Stability
Abstract
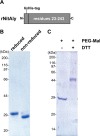
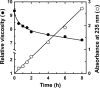
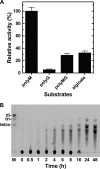
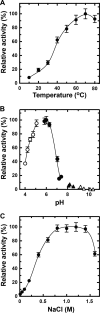
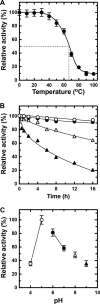
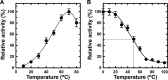
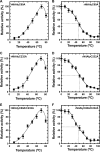
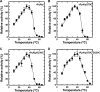
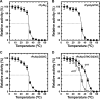
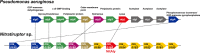
Extremophiles are expected to represent a source of enzymes having unique functional properties. The hypothetical protein NIS_0185, termed NitAly in this study, was identified as an alginate lyase-homolog protein in the genomic database of ϵ-Proteobacteria Nitratiruptor sp. SB155-2, which was isolated from deep-sea hydrothermal vents at a water depth of 1,000 m. Among the characterized alginate lyases in the polysaccharide lyase family 7 (PL-7), the amino acid sequence of NitAly showed the highest identity (39%) with that of red alga Pyropia yezoensis alginate lyase PyAly. Recombinant NitAly (rNitAly) was successfully expressed in Escherichia coli Purified rNitAly degraded alginate in an endolytic manner. Among alginate block types, polyM was preferable to polyG and polyMG as a substrate, and its end degradation products were mainly tri-, tetra-, and penta-saccharides. The optimum temperature and pH values were 70 °C and around 6, respectively. A high concentration of NaCl (0.8-1.4 m) was required for maximum activity. In addition, a 50% loss of activity was observed after incubation at 67 °C for 30 min. Heat stability was decreased in the presence of 5 mm DTT, and Cys-80 and Cys-232 were identified as the residues responsible for heat stability but not lyase activity. Introducing two cysteines into PyAly based on homology modeling using Pseudomonas aeruginosa alginate lyase PA1167 as the template enhanced its heat stability. Thus, NitAly is a functional alginate lyase, with its unique optimum conditions adapted to its environment. These insights into the heat stability of NitAly could be applied to improve that of other PL-7 alginate lyases.
Keywords: algae; alginate; alginate lyase; bacteria; brown algae; carbohydrate; carbohydrate processing; extremophile; heat stability.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures


References
-
- Pawar S. N., and Edgar K. J. (2012) Alginate derivatization: a review of chemistry, properties and applications. Biomaterials 33, 3279–3305 - PubMed
-
- Remminghorst U., and Rehm B. H. (2006) Bacterial alginates: from biosynthesis to applications. Biotechnol. Lett. 28, 1701–1712 - PubMed
-
- Michel G., Tonon T., Scornet D., Cock J. M., and Kloareg B. (2010) The cell wall polysaccharide metabolism of the brown alga Ectocarpus siliculosus. Insights into the evolution of extracellular matrix polysaccharides in eukaryotes. New Phytol. 188, 82–97 - PubMed
-
- Boyd A., and Chakrabarty D. A. (1995) Pseudomonas aeruginosa biofilms: role of the alginate exopolysaccharide. J. Ind. Microbiol. 15, 162–168 - PubMed
-
- Wong T. Y., Preston L. A., and Schiller N. L. (2000) Alginate lyase: review of major sources and enzyme characteristics, structure-function analysis, biological roles, and applications. Annu. Rev. Microbiol. 54, 289–340 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

