In Vivo Tracking of Human Hematopoiesis Reveals Patterns of Clonal Dynamics during Early and Steady-State Reconstitution Phases
- PMID: 27237736
- PMCID: PMC4942697
- DOI: 10.1016/j.stem.2016.04.016
In Vivo Tracking of Human Hematopoiesis Reveals Patterns of Clonal Dynamics during Early and Steady-State Reconstitution Phases
Abstract
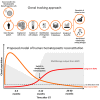
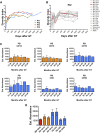
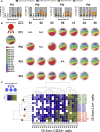
Hematopoietic stem/progenitor cells (HSPCs) are capable of supporting the lifelong production of blood cells exerting a wide spectrum of functions. Lentiviral vector HSPC gene therapy generates a human hematopoietic system stably marked at the clonal level by vector integration sites (ISs). Using IS analysis, we longitudinally tracked >89,000 clones from 15 distinct bone marrow and peripheral blood lineages purified up to 4 years after transplant in four Wiskott-Aldrich syndrome patients treated with HSPC gene therapy. We measured at the clonal level repopulating waves, populations' sizes and dynamics, activity of distinct HSPC subtypes, contribution of various progenitor classes during the early and late post-transplant phases, and hierarchical relationships among lineages. We discovered that in-vitro-manipulated HSPCs retain the ability to return to latency after transplant and can be physiologically reactivated, sustaining a stable hematopoietic output. This study constitutes in vivo comprehensive tracking in humans of hematopoietic clonal dynamics during the early and late post-transplant phases.
Copyright © 2016 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures


References
-
- Aiuti A., Cattaneo F., Galimberti S., Benninghoff U., Cassani B., Callegaro L., Scaramuzza S., Andolfi G., Mirolo M., Brigida I. Gene therapy for immunodeficiency due to adenosine deaminase deficiency. N. Engl. J. Med. 2009;360:447–458. - PubMed
-
- Akashi K., Traver D., Miyamoto T. A clonogenic common myeloid progenitor that gives rise to all myeloid lineages. Nature. 2000;404:193–197. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

