Two shikimate dehydrogenases, VvSDH3 and VvSDH4, are involved in gallic acid biosynthesis in grapevine
- PMID: 27241494
- PMCID: PMC4892741
- DOI: 10.1093/jxb/erw184
Two shikimate dehydrogenases, VvSDH3 and VvSDH4, are involved in gallic acid biosynthesis in grapevine
Abstract
In plants, the shikimate pathway provides aromatic amino acids that are used to generate numerous secondary metabolites, including phenolic compounds. In this pathway, shikimate dehydrogenases (SDH) 'classically' catalyse the reversible dehydrogenation of 3-dehydroshikimate to shikimate. The capacity of SDH to produce gallic acid from shikimate pathway metabolites has not been studied in depth. In grapevine berries, gallic acid mainly accumulates as galloylated flavan-3-ols. The four grapevine SDH proteins have been produced in Escherichia coli In vitro, VvSDH1 exhibited the highest 'classical' SDH activity. Two genes, VvSDH3 and VvSDH4, mainly expressed in immature berry tissues in which galloylated flavan-3-ols are accumulated, encoded enzymes with lower 'classical' activity but were able to produce gallic acid in vitro The over-expression of VvSDH3 in hairy-roots increased the content of aromatic amino acids and hydroxycinnamates, but had little or no effect on molecules more distant from the shikimate pathway (stilbenoids and flavan-3-ols). In parallel, the contents of gallic acid, β-glucogallin, and galloylated flavan-3-ols were increased, attesting to the influence of this gene on gallic acid metabolism. Phylogenetic analysis from dicotyledon SDHs opens the way for the examination of genes from other plants which accumulate gallic acid-based metabolites.
Keywords: Flavan-3-ol; flavonoid; gallic acid; galloylation; grapevine; shikimate dehydrogenase..
© The Author 2016. Published by Oxford University Press on behalf of the Society for Experimental Biology.
Figures







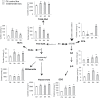
References
-
- Akagi T, Ikegami A, Suzuki Y, Yoshida J, Yamada M, Sato A, Yonemori K. 2009. Expression balances of structural genes in shikimate and flavonoid biosynthesis cause a difference in proanthocyanidin accumulation in persimmon (Diospyros kaki Thunb.) fruit. Planta 230, 899–915. - PubMed
-
- Arnold RA, Noble AC, Singleton VL. 1980. Bitterness and astringency of phenolic fractions in wine. Journal of Agricultural and Food Chemistry 28, 675–678.
-
- Bontpart T, Cheynier V, Ageorges A, Terrier N. 2015. BAHD or SCPL acyltransferase? What a dilemma for acylation in the world of plant phenolic compounds. New Phytologist 208, 695–707. - PubMed
-
- Boulekbache-Makhlouf L, Meudec E, Chibane M, Mazauric JP, Slimani S, Henry M, Cheynier V, Madani K. 2010. Analysis by high-performance liquid chromatography diode array detection mass spectrometry of phenolic compounds in fruit of Eucalyptus globulus cultivated in Algeria. Journal of Agricultural and Food Chemistry 58, 12615–24. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

