Human T cell responses to Japanese encephalitis virus in health and disease
- PMID: 27242166
- PMCID: PMC4925015
- DOI: 10.1084/jem.20151517
Human T cell responses to Japanese encephalitis virus in health and disease
Abstract
Japanese encephalitis (JE) virus (JEV) is an important cause of encephalitis in children of South and Southeast Asia. However, the majority of individuals exposed to JEV only develop mild symptoms associated with long-lasting adaptive immunity. The related flavivirus dengue virus (DENV) cocirculates in many JEV-endemic areas, and clinical data suggest cross-protection between DENV and JEV. To address the role of T cell responses in protection against JEV, we conducted the first full-breadth analysis of the human memory T cell response using a synthetic peptide library. Ex vivo interferon-γ (IFN-γ) responses to JEV in healthy JEV-exposed donors were mostly CD8(+) and targeted nonstructural (NS) proteins, whereas IFN-γ responses in recovered JE patients were mostly CD4(+) and targeted structural proteins and the secreted protein NS1. Among patients, a high quality, polyfunctional CD4(+) T cell response was associated with complete recovery from JE. T cell responses from healthy donors showed a high degree of cross-reactivity to DENV that was less apparent in recovered JE patients despite equal exposure. These data reveal divergent functional CD4(+) and CD8(+) T cell responses linked to different clinical outcomes of JEV infection, associated with distinct targeting and broad flavivirus cross-reactivity including epitopes from DENV, West Nile, and Zika virus.
© 2016 Turtle et al.
Figures

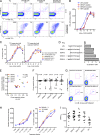
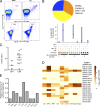



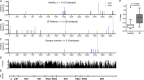
References
-
- Akondy R.S., Monson N.D., Miller J.D., Edupuganti S., Teuwen D., Wu H., Quyyumi F., Garg S., Altman J.D., Del Rio C., et al. . 2009. The yellow fever virus vaccine induces a broad and polyfunctional human memory CD8+ T cell response. J. Immunol. 183:7919–7930. 10.4049/jimmunol.0803903 - DOI - PMC - PubMed
-
- Anderson K.B., Gibbons R.V., Thomas S.J., Rothman A.L., Nisalak A., Berkelman R.L., Libraty D.H., and Endy T.P.. 2011. Preexisting Japanese encephalitis virus neutralizing antibodies and increased symptomatic dengue illness in a school-based cohort in Thailand. PLoS Negl. Trop. Dis. 5:e1311 10.1371/journal.pntd.0001311 - DOI - PMC - PubMed
-
- Appanna R., Huat T.L., See L.L., Tan P.L., Vadivelu J., and Devi S.. 2007. Cross-reactive T-cell responses to the nonstructural regions of dengue viruses among dengue fever and dengue hemorrhagic fever patients in Malaysia. Clin. Vaccine Immunol. 14:969–977. 10.1128/CVI.00069-07 - DOI - PMC - PubMed
-
- Ashok M.S., and Rangarajan P.N.. 1999. Immunization with plasmid DNA encoding the envelope glycoprotein of Japanese Encephalitis virus confers significant protection against intracerebral viral challenge without inducing detectable antiviral antibodies. Vaccine. 18:68–75. 10.1016/S0264-410X(99)00180-2 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

