Sunflower Resistance to Broomrape (Orobanche cumana) Is Controlled by Specific QTLs for Different Parasitism Stages
- PMID: 27242810
- PMCID: PMC4861731
- DOI: 10.3389/fpls.2016.00590
Sunflower Resistance to Broomrape (Orobanche cumana) Is Controlled by Specific QTLs for Different Parasitism Stages
Abstract
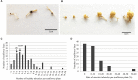
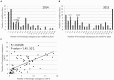
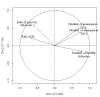
Orobanche cumana (sunflower broomrape) is an obligatory and non-photosynthetic root parasitic plant that specifically infects the sunflower. It is located in Europe and in Asia, where it can cause yield losses of over 80%. More aggressive races have evolved, mainly around the Black Sea, and broomrape can rapidly spread to new areas. Breeding for resistance seems to be the most efficient and sustainable approach to control broomrape infestation. In our study, we used a population of 101 recombinant inbred lines (RILs), derived from a cross between the two lines HA89 and LR1 (a line derived from an interspecific cross with Helianthus debilis). Rhizotrons, pots and field experiments were used to characterize all RILs for their resistance to O. cumana race F parasitism at three post vascular connection life stages: (i) early attachment of the parasite to the sunflower roots, (ii) young tubercle and (iii) shoot emergence. In addition, RIL resistance to race G at young tubercle development stage was evaluated in pots. The entire population was genotyped, and QTLs were mapped. Different QTLs were identified for each race (F from Spain and G from Turkey) and for the three stages of broomrape development. The results indicate that there are several quantitative resistance mechanisms controlling the infection by O. cumana that can be used in sunflower breeding.
Keywords: Orobanche cumana; QTL mapping; broomrape; candidate genes; parasitic weeds; plant–plant interaction; resistance; sunflower.
Figures




Similar articles
-
Quantitative trait loci for broomrape (Orobanche cumana Wallr.) resistance in sunflower.Theor Appl Genet. 2004 Jun;109(1):92-102. doi: 10.1007/s00122-004-1599-7. Epub 2004 Feb 13. Theor Appl Genet. 2004. PMID: 14968309
-
Development and characterization of a new sunflower source of resistance to race G of Orobanche cumana Wallr. derived from Helianthus anomalus.Theor Appl Genet. 2024 Feb 22;137(3):56. doi: 10.1007/s00122-024-04558-4. Theor Appl Genet. 2024. PMID: 38386181 Free PMC article.
-
First Report of Race Composition and Distribution of Sunflower Broomrape, Orobanche cumana, in China.Plant Dis. 2015 Feb;99(2):291. doi: 10.1094/PDIS-07-14-0721-PDN. Plant Dis. 2015. PMID: 30699572
-
Genetic and Genomic Tools in Sunflower Breeding for Broomrape Resistance.Genes (Basel). 2020 Jan 30;11(2):152. doi: 10.3390/genes11020152. Genes (Basel). 2020. PMID: 32019223 Free PMC article. Review.
-
Broomrape Weeds. Underground Mechanisms of Parasitism and Associated Strategies for their Control: A Review.Front Plant Sci. 2016 Feb 19;7:135. doi: 10.3389/fpls.2016.00135. eCollection 2016. Front Plant Sci. 2016. PMID: 26925071 Free PMC article. Review.
Cited by
-
Green synthesis and characterization of Fe2O3, ZnO and TiO2 nanoparticles and searching for their potential use as biofertilizer on sunflower.Physiol Mol Biol Plants. 2024 Sep;30(9):1429-1447. doi: 10.1007/s12298-024-01508-8. Epub 2024 Sep 12. Physiol Mol Biol Plants. 2024. PMID: 39310700
-
Sunflower Hybrid Breeding: From Markers to Genomic Selection.Front Plant Sci. 2018 Jan 17;8:2238. doi: 10.3389/fpls.2017.02238. eCollection 2017. Front Plant Sci. 2018. PMID: 29387071 Free PMC article. Review.
-
Response of faba bean to intercropping, biological and chemical control against broomrape and root rot diseases.Saudi J Biol Sci. 2022 May;29(5):3482-3493. doi: 10.1016/j.sjbs.2022.02.032. Epub 2022 Feb 25. Saudi J Biol Sci. 2022. PMID: 35844392 Free PMC article.
-
Different genetic architectures underlie crop responses to the same pathogen: the {Helianthus annuus * Phoma macdonaldii} interaction case for black stem disease and premature ripening.BMC Plant Biol. 2017 Oct 19;17(1):167. doi: 10.1186/s12870-017-1116-1. BMC Plant Biol. 2017. PMID: 29052528 Free PMC article.
-
Aspergillus alliaceus infection fatally shifts Orobanche hormones and phenolic metabolism.Braz J Microbiol. 2020 Sep;51(3):883-892. doi: 10.1007/s42770-020-00283-4. Epub 2020 May 3. Braz J Microbiol. 2020. PMID: 32363566 Free PMC article.
References
-
- Alcántara E., Morales-García M., Díaz-Sánchez J. (2006). Effects of broomrape parasitism on sunflower plants: growth, development, and mineral nutrition. J. Plant Nutr. 29 1199–1206. 10.1080/01904160600767351 - DOI
-
- Aly R., Cholakh H., Joel D. M., Leibman D., Steinitz B., Zelcer A., et al. (2009). Gene silencing of mannose 6-phosphate reductase in the parasitic weed Orobanche aegyptiaca through the production of homologous dsRNA sequences in the host plant. Plant Biotechnol. J. 7 487–498. 10.1111/j.1467-7652.2009.00418.x - DOI - PubMed
-
- Clark R. T., Famoso A. N., Zhao K., Shaff J. E., Craft E. J., Bustamante C. D., et al. (2013). High-throughput two-dimensional root system phenotyping platform facilitates genetic analysis of root growth and development: root phenotyping platform. Plant Cell Environ. 36 454–466. 10.1111/j.1365-3040.2012.02587.x - DOI - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

