Human Galectin-9 Is a Potent Mediator of HIV Transcription and Reactivation
- PMID: 27253379
- PMCID: PMC4890776
- DOI: 10.1371/journal.ppat.1005677
Human Galectin-9 Is a Potent Mediator of HIV Transcription and Reactivation
Abstract
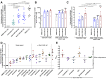
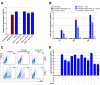
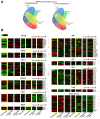
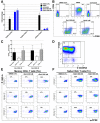
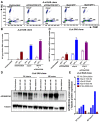
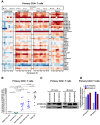
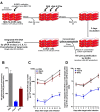
Identifying host immune determinants governing HIV transcription, latency and infectivity in vivo is critical to developing an HIV cure. Based on our recent finding that the host factor p21 regulates HIV transcription during antiretroviral therapy (ART), and published data demonstrating that the human carbohydrate-binding immunomodulatory protein galectin-9 regulates p21, we hypothesized that galectin-9 modulates HIV transcription. We report that the administration of a recombinant, stable form of galectin-9 (rGal-9) potently reverses HIV latency in vitro in the J-Lat HIV latency model. Furthermore, rGal-9 reverses HIV latency ex vivo in primary CD4+ T cells from HIV-infected, ART-suppressed individuals (p = 0.002), more potently than vorinostat (p = 0.02). rGal-9 co-administration with the latency reversal agent "JQ1", a bromodomain inhibitor, exhibits synergistic activity (p<0.05). rGal-9 signals through N-linked oligosaccharides and O-linked hexasaccharides on the T cell surface, modulating the gene expression levels of key transcription initiation, promoter proximal-pausing, and chromatin remodeling factors that regulate HIV latency. Beyond latent viral reactivation, rGal-9 induces robust expression of the host antiviral deaminase APOBEC3G in vitro and ex vivo (FDR<0.006) and significantly reduces infectivity of progeny virus, decreasing the probability that the HIV reservoir will be replenished when latency is reversed therapeutically. Lastly, endogenous levels of soluble galectin-9 in the plasma of 72 HIV-infected ART-suppressed individuals were associated with levels of HIV RNA in CD4+ T cells (p<0.02) and with the quantity and binding avidity of circulating anti-HIV antibodies (p<0.009), suggesting a role of galectin-9 in regulating HIV transcription and viral production in vivo during therapy. Our data suggest that galectin-9 and the host glycosylation machinery should be explored as foundations for novel HIV cure strategies.
Conflict of interest statement
I have read the journal's policy and the authors of this manuscript have the following competing interests: MLS is currently employed by RainDance Technologies, Inc., which provided support for some of the droplet digital PCR assays performed in this study to measure HIV and host gene expression. TN and MH are employed by GalPharma Co., Ltd., which designed and provided the recombinant, stable form galectin-9 used in our study, and supported experiments to evaluate the safety and tolerability of recombinant galactic-9 in the Lewis rat model. These relationships do not alter our adherence to all PLoS Pathogens policies on sharing data and materials.
Figures


References
-
- Wong JK, Hezareh M, Gunthard HF, Havlir DV, Ignacio CC, et al. (1997) Recovery of replication-competent HIV despite prolonged suppression of plasma viremia. Science 278: 1291–1295. - PubMed
-
- Finzi D, Hermankova M, Pierson T, Carruth LM, Buck C, et al. (1997) Identification of a reservoir for HIV-1 in patients on highly active antiretroviral therapy. Science 278: 1295–1300. - PubMed
-
- Saez-Cirion A, Bacchus C, Hocqueloux L, Avettand-Fenoel V, Girault I, et al. (2013) Post-treatment HIV-1 controllers with a long-term virological remission after the interruption of early initiated antiretroviral therapy ANRS VISCONTI Study. PLoS Pathog 9: e1003211 10.1371/journal.ppat.1003211 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

