Chemical genetic discovery of PARP targets reveals a role for PARP-1 in transcription elongation
- PMID: 27256882
- PMCID: PMC5540732
- DOI: 10.1126/science.aaf7865
Chemical genetic discovery of PARP targets reveals a role for PARP-1 in transcription elongation
Abstract
Poly[adenosine diphosphate (ADP)-ribose] polymerases (PARPs) are a family of enzymes that modulate diverse biological processes through covalent transfer of ADP-ribose from the oxidized form of nicotinamide adenine dinucleotide (NAD(+)) onto substrate proteins. Here we report a robust NAD(+) analog-sensitive approach for PARPs, which allows PARP-specific ADP-ribosylation of substrates that is suitable for subsequent copper-catalyzed azide-alkyne cycloaddition reactions. Using this approach, we mapped hundreds of sites of ADP-ribosylation for PARPs 1, 2, and 3 across the proteome, as well as thousands of PARP-1-mediated ADP-ribosylation sites across the genome. We found that PARP-1 ADP-ribosylates and inhibits negative elongation factor (NELF), a protein complex that regulates promoter-proximal pausing by RNA polymerase II (Pol II). Depletion or inhibition of PARP-1 or mutation of the ADP-ribosylation sites on NELF-E promotes Pol II pausing, providing a clear functional link between PARP-1, ADP-ribosylation, and NELF. This analog-sensitive approach should be broadly applicable across the PARP family and has the potential to illuminate the ADP-ribosylated proteome and the molecular mechanisms used by individual PARPs to mediate their responses to cellular signals.
Copyright © 2016, American Association for the Advancement of Science.
Figures
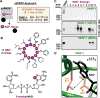





Comment in
-
Post-translational modifications: ADP-ribosylation promotes transcription elongation.Nat Rev Mol Cell Biol. 2016 Jun 22;17(7):397. doi: 10.1038/nrm.2016.83. Nat Rev Mol Cell Biol. 2016. PMID: 27329964 No abstract available.
-
Identification of PARP-Specific ADP-Ribosylation Targets Reveals a Regulatory Function for ADP-Ribosylation in Transcription Elongation.Mol Cell. 2016 Jul 21;63(2):181-183. doi: 10.1016/j.molcel.2016.07.006. Mol Cell. 2016. PMID: 27447982
References
-
- Gibson BA, Kraus WL. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat Rev Mol Cell Biol. 2012;13:411. - PubMed
-
- Hottiger MO. Nuclear ADP-ribosylation and its role in chromatin plasticity, cell differentiation, and epigenetics. Annu Rev Biochem. 2015;84:227. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

