Interplay between Metabolism and Epigenetics: A Nuclear Adaptation to Environmental Changes
- PMID: 27259202
- PMCID: PMC4893201
- DOI: 10.1016/j.molcel.2016.05.029
Interplay between Metabolism and Epigenetics: A Nuclear Adaptation to Environmental Changes
Abstract
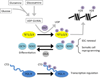
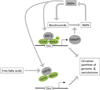
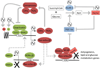
The physiological identity of every cell is maintained by highly specific transcriptional networks that establish a coherent molecular program that is in tune with nutritional conditions. The regulation of cell-specific transcriptional networks is accomplished by an epigenetic program via chromatin-modifying enzymes, whose activity is directly dependent on metabolites such as acetyl-coenzyme A, S-adenosylmethionine, and NAD+, among others. Therefore, these nuclear activities are directly influenced by the nutritional status of the cell. In addition to nutritional availability, this highly collaborative program between epigenetic dynamics and metabolism is further interconnected with other environmental cues provided by the day-night cycles imposed by circadian rhythms. Herein, we review molecular pathways and their metabolites associated with epigenetic adaptations modulated by histone- and DNA-modifying enzymes and their responsiveness to the environment in the context of health and disease.
Copyright © 2016 Elsevier Inc. All rights reserved.
Figures


References
-
- Akhtar RA, Reddy AB, Maywood ES, Clayton JD, King VM, Smith AG, Gant TW, Hastings MH, Kyriacou CP. Circadian cycling of the mouse liver transcriptome, as revealed by cDNA microarray, is driven by the suprachiasmatic nucleus. Curr Biol. 2002;12:540–550. - PubMed
-
- Asher G, Gatfield D, Stratmann M, Reinke H, Dibner C, Kreppel F, Mostoslavsky R, Alt FW, Schibler U. SIRT1 regulates circadian clock gene expression through PER2 deacetylation. Cell. 2008;134:317–328. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

