When DNA repair goes wrong: BER-generated DNA-protein crosslinks to oxidative lesions
- PMID: 27264558
- PMCID: PMC6420214
- DOI: 10.1016/j.dnarep.2016.05.014
When DNA repair goes wrong: BER-generated DNA-protein crosslinks to oxidative lesions
Abstract
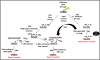
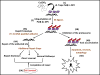
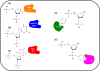
Free radicals generate an array of DNA lesions affecting all parts of the molecule. The damage to deoxyribose receives less attention than base damage, even though the former accounts for ∼20% of the total. Oxidative deoxyribose fragments (e.g., 3'-phosphoglycolate esters) are removed by the Ape1 AP endonuclease and other enzymes in mammalian cells to enable DNA repair synthesis. Oxidized abasic sites are initially incised by Ape1, thus recruiting these lesions into base excision repair (BER) pathways. Lesions such as 2-deoxypentos-4-ulose can be removed by conventional (single-nucleotide) BER, which proceeds through a covalent Schiff base intermediate with DNA polymerase β (Polβ) that is resolved by hydrolysis. In contrast, the lesion 2-deoxyribonolactone (dL) must be processed by multinucleotide ("long-patch") BER: attempted repair via the single-nucleotide pathway leads to a dead-end, covalent complex with Polβ cross- linked to the DNA by an amide bond. We recently detected these stable DNA-protein crosslinks (DPC) between Polβ and dL in intact cells. The features of the DPC formation in vivo are exactly in keeping with the mechanistic properties seen in vitro: Polβ-DPC are formed by oxidative agents in line with their ability to form the dL lesion; they are not formed by non-oxidative agents; DPC formation absolutely requires the active-site lysine-72 that attacks the 5'-deoxyribose; and DPC formation depends on Ape1 to incise the dL lesion first. The Polβ-DPC are rapidly processed in vivo, the signal disappearing with a half-life of 15-30min in both mouse and human cells. This removal is blocked by inhibiting the proteasome, which leads to the accumulation of ubiquitin associated with the Polβ-DPC. While other proteins (e.g., topoisomerases) also form DPC under these conditions, 60-70% of the trapped ubiquitin depends on Polβ. The mechanism of ubiquitin targeting to Polβ-DPC, the subsequent processing of the expected 5'-peptidyl-dL, and the biological consequences of unrepaired DPC are important to assess. Many other lyase enzymes that attack dL can also be trapped in DPC, so the processing mechanisms may apply quite broadly.
Keywords: 2-deoxyribonolactone; AP lyase; Base excision DNA repair; DNA polymerase β; DNA-protein crosslinks; Oxidative DNA damage.
Copyright © 2016 Elsevier B.V. All rights reserved.
Figures
Similar articles
-
Oxidative DNA-protein crosslinks formed in mammalian cells by abasic site lyases involved in DNA repair.DNA Repair (Amst). 2020 Mar;87:102773. doi: 10.1016/j.dnarep.2019.102773. Epub 2020 Jan 9. DNA Repair (Amst). 2020. PMID: 31945542 Free PMC article.
-
C-terminal residues of DNA polymerase β and E3 ligase required for ubiquitin-linked proteolysis of oxidative DNA-protein crosslinks.DNA Repair (Amst). 2024 Nov;143:103756. doi: 10.1016/j.dnarep.2024.103756. Epub 2024 Aug 28. DNA Repair (Amst). 2024. PMID: 39243487
-
Enzyme mechanism-based, oxidative DNA-protein cross-links formed with DNA polymerase β in vivo.Proc Natl Acad Sci U S A. 2015 Jul 14;112(28):8602-7. doi: 10.1073/pnas.1501101112. Epub 2015 Jun 29. Proc Natl Acad Sci U S A. 2015. PMID: 26124145 Free PMC article.
-
Roles of base excision repair subpathways in correcting oxidized abasic sites in DNA.FEBS J. 2006 Apr;273(8):1620-9. doi: 10.1111/j.1742-4658.2006.05192.x. FEBS J. 2006. PMID: 16623699 Review.
-
Molecular and biological roles of Ape1 protein in mammalian base excision repair.DNA Repair (Amst). 2005 Dec 8;4(12):1442-9. doi: 10.1016/j.dnarep.2005.09.004. Epub 2005 Sep 30. DNA Repair (Amst). 2005. PMID: 16199212 Review.
Cited by
-
RNA polymerase pausing, stalling and bypass during transcription of damaged DNA: from molecular basis to functional consequences.Nucleic Acids Res. 2022 Apr 8;50(6):3018-3041. doi: 10.1093/nar/gkac174. Nucleic Acids Res. 2022. PMID: 35323981 Free PMC article. Review.
-
Risky repair: DNA-protein crosslinks formed by mitochondrial base excision DNA repair enzymes acting on free radical lesions.Free Radic Biol Med. 2017 Jun;107:146-150. doi: 10.1016/j.freeradbiomed.2016.11.025. Epub 2016 Nov 17. Free Radic Biol Med. 2017. PMID: 27867099 Free PMC article. Review.
-
A quantitative PCR-based assay reveals that nucleotide excision repair plays a predominant role in the removal of DNA-protein crosslinks from plasmids transfected into mammalian cells.DNA Repair (Amst). 2018 Feb;62:18-27. doi: 10.1016/j.dnarep.2018.01.004. Epub 2018 Jan 9. DNA Repair (Amst). 2018. PMID: 29413806 Free PMC article.
-
Base Excision Repair: Mechanisms and Impact in Biology, Disease, and Medicine.Int J Mol Sci. 2023 Sep 16;24(18):14186. doi: 10.3390/ijms241814186. Int J Mol Sci. 2023. PMID: 37762489 Free PMC article. Review.
-
New insights into abasic site repair and tolerance.DNA Repair (Amst). 2020 Jun;90:102866. doi: 10.1016/j.dnarep.2020.102866. Epub 2020 Apr 30. DNA Repair (Amst). 2020. PMID: 32417669 Free PMC article. Review.
References
-
- Nakamura J, Swenberg JA. Endogenous Apurinic/Apyrimidinic Sites in Genomic DNA of Mammalian Tissues. Cancer Research. 1999;59(11):2522–6. - PubMed
-
- Nakamura J, Walker VE, Upton PB, Chiang S-Y, Kow YW, Swenberg JA. Highly Sensitive Apurinic/Apyrimidinic Site Assay Can Detect Spontaneous and Chemically Induced Depurination under Physiological Conditions. Cancer Research. 1998;58(2):222–5. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

