Fast and accurate long-range phasing in a UK Biobank cohort
- PMID: 27270109
- PMCID: PMC4925291
- DOI: 10.1038/ng.3571
Fast and accurate long-range phasing in a UK Biobank cohort
Abstract
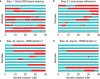
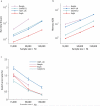
Recent work has leveraged the extensive genotyping of the Icelandic population to perform long-range phasing (LRP), enabling accurate imputation and association analysis of rare variants in target samples typed on genotyping arrays. Here we develop a fast and accurate LRP method, Eagle, that extends this paradigm to populations with much smaller proportions of genotyped samples by harnessing long (>4-cM) identical-by-descent (IBD) tracts shared among distantly related individuals. We applied Eagle to N ≈ 150,000 samples (0.2% of the British population) from the UK Biobank, and we determined that it is 1-2 orders of magnitude faster than existing methods while achieving similar or better phasing accuracy (switch error rate ≈ 0.3%, corresponding to perfect phase in a majority of 10-Mb segments). We also observed that, when used within an imputation pipeline, Eagle prephasing improved downstream imputation accuracy in comparison to prephasing in batches using existing methods, as necessary to achieve comparable computational cost.
Figures
References
-
- Marchini J, Howie B, Myers S, McVean G, Donnelly P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nature Genetics. 2007;39:906–913. - PubMed
-
- Marchini J, Howie B. Genotype imputation for genome-wide association studies. Nature Reviews Genetics. 2010;11:499–511. - PubMed
References (Online Methods)
-
- Indyk P, Motwani R. Proceedings of the thirtieth annual ACM Symposium on Theory of Computing. ACM; 1998. Approximate nearest neighbors: towards removing the curse of dimensionality. pp. 604–613.
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

