Targeted gene silencing of CCL2 inhibits triple negative breast cancer progression by blocking cancer stem cell renewal and M2 macrophage recruitment
- PMID: 27283985
- PMCID: PMC5226513
- DOI: 10.18632/oncotarget.9885
Targeted gene silencing of CCL2 inhibits triple negative breast cancer progression by blocking cancer stem cell renewal and M2 macrophage recruitment
Abstract
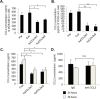
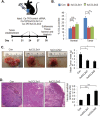
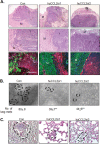
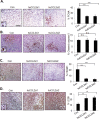
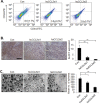
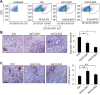
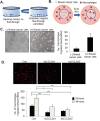
Triple negative breast cancers are an aggressive subtype of breast cancer, characterized by the lack of estrogen receptor, progesterone receptor and Her2 expression. Triple negative breast cancers are non-responsive to conventional anti-hormonal and Her2 targeted therapies, making it necessary to identify new molecular targets for therapy. The chemokine CCL2 is overexpressed in invasive breast cancers, and regulates breast cancer progression through multiple mechanisms. With few approaches to target CCL2 activity, its value as a therapeutic target is unclear. In these studies, we developed a novel gene silencing approach that involves complexing siRNAs to TAT cell penetrating peptides (Ca-TAT) through non-covalent calcium cross-linking. Ca-TAT/siRNA complexes penetrated 3D collagen cultures of breast cancer cells and inhibited CCL2 expression more effectively than conventional antibody neutralization. Ca-TAT/siRNA complexes targeting CCL2 were delivered to mice bearing MDA-MB-231 breast tumor xenografts. In vivo CCL2 gene silencing inhibited primary tumor growth and metastasis, associated with a reduction in cancer stem cell renewal and recruitment of M2 macrophages. These studies are the first to demonstrate that targeting CCL2 expression in vivo may be a viable therapeutic approach to treating triple negative breast cancer.
Keywords: CCL2; TAT cell penetrating peptide; breast cancer; cancer stem cell; macrophage.
Conflict of interest statement
N. Cheng and W. Fang were scientific advisors to Meta Bioscience LLC.
Figures
Similar articles
-
Hesperidin PLGA nanoparticles potentiate the efficacy of aPD-1 in treating triple negative breast cancer by regulating CCL2 and ADPN expression in cancer-associated adipocytes.Int Immunopharmacol. 2024 Oct 25;140:112759. doi: 10.1016/j.intimp.2024.112759. Epub 2024 Aug 3. Int Immunopharmacol. 2024. PMID: 39098226
-
Tumor-associated macrophages promote epithelial-mesenchymal transition and the cancer stem cell properties in triple-negative breast cancer through CCL2/AKT/β-catenin signaling.Cell Commun Signal. 2022 Jun 17;20(1):92. doi: 10.1186/s12964-022-00888-2. Cell Commun Signal. 2022. PMID: 35715860 Free PMC article.
-
CCR2 Chemokine Receptors Enhance Growth and Cell-Cycle Progression of Breast Cancer Cells through SRC and PKC Activation.Mol Cancer Res. 2019 Feb;17(2):604-617. doi: 10.1158/1541-7786.MCR-18-0750. Epub 2018 Nov 16. Mol Cancer Res. 2019. PMID: 30446625 Free PMC article.
-
MCT-1/miR-34a/IL-6/IL-6R signaling axis promotes EMT progression, cancer stemness and M2 macrophage polarization in triple-negative breast cancer.Mol Cancer. 2019 Mar 18;18(1):42. doi: 10.1186/s12943-019-0988-0. Mol Cancer. 2019. PMID: 30885232 Free PMC article.
-
The CCL2 chemokine is a negative regulator of autophagy and necrosis in luminal B breast cancer cells.Breast Cancer Res Treat. 2015 Apr;150(2):309-20. doi: 10.1007/s10549-015-3324-4. Epub 2015 Mar 6. Breast Cancer Res Treat. 2015. PMID: 25744294 Free PMC article.
Cited by
-
The multifaceted roles of the chemokines CCL2 and CXCL12 in osteophilic metastatic cancers.Cancer Metastasis Rev. 2021 Jun;40(2):427-445. doi: 10.1007/s10555-021-09974-2. Epub 2021 May 11. Cancer Metastasis Rev. 2021. PMID: 33973098 Review.
-
Breast Cancer Stem Cells: Signaling Pathways, Cellular Interactions, and Therapeutic Implications.Cancers (Basel). 2022 Jul 5;14(13):3287. doi: 10.3390/cancers14133287. Cancers (Basel). 2022. PMID: 35805056 Free PMC article. Review.
-
Fundamentals of siRNA and miRNA therapeutics and a review of targeted nanoparticle delivery systems in breast cancer.Biophys Rev. 2018 Feb;10(1):69-86. doi: 10.1007/s12551-017-0392-1. Epub 2018 Jan 11. Biophys Rev. 2018. PMID: 29327101 Free PMC article. Review.
-
Eukaryotic elongation factor-2 kinase (eEF2K) signaling in tumor and microenvironment as a novel molecular target.J Mol Med (Berl). 2020 Jun;98(6):775-787. doi: 10.1007/s00109-020-01917-8. Epub 2020 May 7. J Mol Med (Berl). 2020. PMID: 32377852 Review.
-
AR negative triple negative or "quadruple negative" breast cancers in African American women have an enriched basal and immune signature.PLoS One. 2018 Jun 18;13(6):e0196909. doi: 10.1371/journal.pone.0196909. eCollection 2018. PLoS One. 2018. PMID: 29912871 Free PMC article. Clinical Trial.
References
-
- Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS, Thorsen T, Quist H, Matese JC, Brown PO, Botstein D, Eystein Lonning P, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A. 2001;98:10869–10874. - PMC - PubMed
-
- Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A, Deng S, Johnsen H, Pesich R, Geisler S, Demeter J, Perou CM, Lonning PE, Brown PO, Borresen-Dale AL, Botstein D. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci U S A. 2003;100:8418–8423. - PMC - PubMed
-
- Nielsen TO, Hsu FD, Jensen K, Cheang M, Karaca G, Hu Z, Hernandez-Boussard T, Livasy C, Cowan D, Dressler L, Akslen LA, Ragaz J, Gown AM, Gilks CB, van de Rijn M, Perou CM. Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res. 2004;10:5367–5374. - PubMed
-
- Livasy CA, Karaca G, Nanda R, Tretiakova MS, Olopade OI, Moore DT, Perou CM. Phenotypic evaluation of the basal-like subtype of invasive breast carcinoma. Modern pathology. 2006;19:264–271. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

