Surprisingly rich repertoire of Wnt genes in the demosponge Halisarca dujardini
- PMID: 27287511
- PMCID: PMC4902976
- DOI: 10.1186/s12862-016-0700-6
Surprisingly rich repertoire of Wnt genes in the demosponge Halisarca dujardini
Abstract
Background: Wnt proteins are secreted signalling molecules found in all animal phyla. In bilaterian animals, including humans, Wnt proteins play key roles in development, maintenance of homeostasis and regeneration. While Wnt gene repertoires and roles are strongly conserved between cnidarians and bilaterians, Wnt genes from basal metazoans (sponges, ctenophores, placozoans) are difficult or impossible to assign to the bilaterian + cnidarian orthologous groups. Moreover, dramatic differences in Wnt numbers among basal metazoan exist, with only three present in the genome of Amphimedon queenslandica, a demosponge, and 21 in the genome of Sycon ciliatum, a calcisponge. To gain insight into the ancestral Wnt repertoire and function, we have chosen to investigate Wnt genes in Halisarca dujardini, a demosponge with relatively well described development and regeneration, and a very distant phylogenetic relationship to Amphimedon.
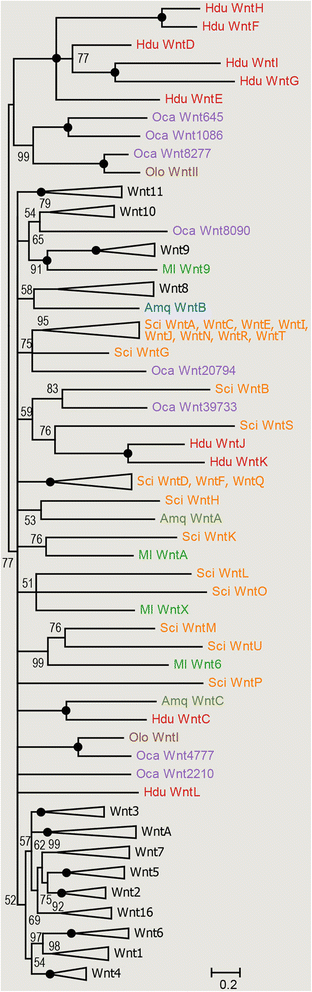
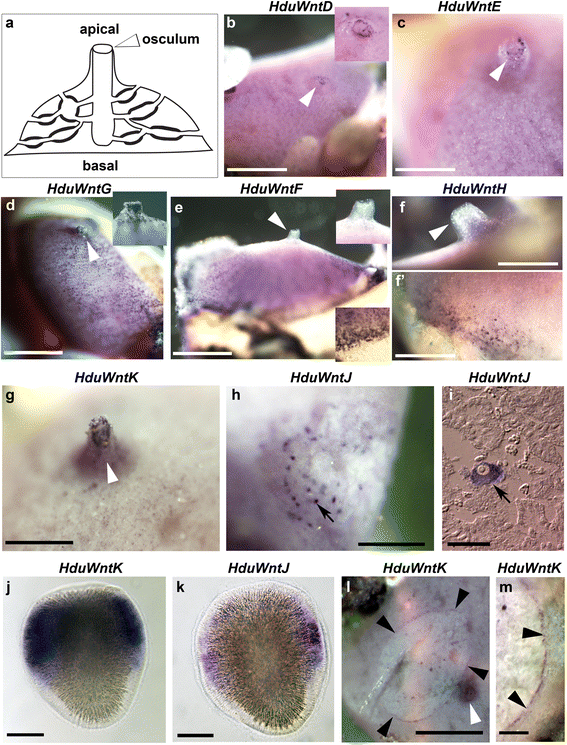
Results: Here we describe generation of a eukaryotic contamination-free transcriptome of Halisarca dujardini, and analysis of Wnt genes repertoire and expression in this species. We have identified ten Wnt genes, with only one orthologous to Amphimedon Wnt, and six appearing to be a result of a lineage specific expansion. Expression analysis carried out by in situ hybridization of adults and larvae revealed that two Halisarca Wnts are expressed in nested domains in the posterior half of the larvae, and six along the adult body axis, with two specific to the osculum. Strikingly, expression of one of the Wnt genes was elevated in the region undergoing regeneration.
Conclusions: Our results demonstrated that the three Poriferan lineages (Demospongiae, Calcarea and Homoloscleromorpha) are characterized by highly diverse Wnt gene repertoires which do not display higher similarity to each other than they do to the non-sponge (i.e. ctenophore, cnidarian and bilaterian) repertoires. This is in striking contrast to the uniform Wnt repertoires in Cnidarians and Bilaterians, suggesting that the Wnt family composition became "fixed" only in the last common ancestor of Cnidarians and Bilaterians. In contrast, expression of Wnt genes in the apical region of sponge adults and the posterior region of sponge larvae suggests conservation of the Wnt role in axial patterning across the animal kingdom.
Figures
Similar articles
-
Structure and expression of conserved Wnt pathway components in the demosponge Amphimedon queenslandica.Evol Dev. 2010 Sep-Oct;12(5):494-518. doi: 10.1111/j.1525-142X.2010.00435.x. Evol Dev. 2010. PMID: 20883218
-
Wnt and TGF-beta expression in the sponge Amphimedon queenslandica and the origin of metazoan embryonic patterning.PLoS One. 2007 Oct 10;2(10):e1031. doi: 10.1371/journal.pone.0001031. PLoS One. 2007. PMID: 17925879 Free PMC article.
-
Comparative analyses of developmental transcription factor repertoires in sponges reveal unexpected complexity of the earliest animals.Mar Genomics. 2015 Dec;24 Pt 2:121-9. doi: 10.1016/j.margen.2015.07.008. Epub 2015 Aug 5. Mar Genomics. 2015. PMID: 26253310 Review.
-
Origin and evolution of laminin gene family diversity.Mol Biol Evol. 2012 Jul;29(7):1823-36. doi: 10.1093/molbev/mss060. Epub 2012 Feb 1. Mol Biol Evol. 2012. PMID: 22319142
-
From traveler to homebody: Which signaling mechanisms sponge larvae use to become adult sponges?Adv Protein Chem Struct Biol. 2019;116:421-449. doi: 10.1016/bs.apcsb.2019.02.002. Epub 2019 Mar 14. Adv Protein Chem Struct Biol. 2019. PMID: 31036299 Review.
Cited by
-
Wnt evolution and function shuffling in liberal and conservative chordate genomes.Genome Biol. 2018 Jul 25;19(1):98. doi: 10.1186/s13059-018-1468-3. Genome Biol. 2018. PMID: 30045756 Free PMC article.
-
Embryonic expression of priapulid Wnt genes.Dev Genes Evol. 2019 Jul;229(4):125-135. doi: 10.1007/s00427-019-00636-6. Epub 2019 Jul 4. Dev Genes Evol. 2019. PMID: 31273439 Free PMC article.
-
Body-Plan Reorganization in a Sponge Correlates with Microbiome Change.Mol Biol Evol. 2023 Jun 1;40(6):msad138. doi: 10.1093/molbev/msad138. Mol Biol Evol. 2023. PMID: 37288516 Free PMC article.
-
Structure of the RECK CC domain, an evolutionary anomaly.Proc Natl Acad Sci U S A. 2020 Jun 30;117(26):15104-15111. doi: 10.1073/pnas.2006332117. Epub 2020 Jun 15. Proc Natl Acad Sci U S A. 2020. PMID: 32541044 Free PMC article.
-
Combinatorial Wnt signaling landscape during brachiopod anteroposterior patterning.BMC Biol. 2024 Sep 19;22(1):212. doi: 10.1186/s12915-024-01988-w. BMC Biol. 2024. PMID: 39300453 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

