Genome-Wide Analysis of Long Noncoding RNAs and Their Responses to Drought Stress in Cotton (Gossypium hirsutum L.)
- PMID: 27294517
- PMCID: PMC4905672
- DOI: 10.1371/journal.pone.0156723
Genome-Wide Analysis of Long Noncoding RNAs and Their Responses to Drought Stress in Cotton (Gossypium hirsutum L.)
Abstract
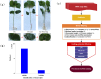
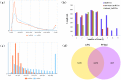
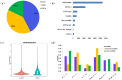
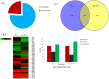
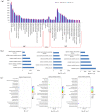
Recent researches on long noncoding RNAs (lncRNAs) have expanded our horizon of gene regulation and the cellular complexity. However, the number, characteristics and expression patterns of lncRNAs remain poorly characterized and how these lncRNAs biogenesis are regulated in response to drought stress in cotton are still largely unclear. In the study, using a reproducibility-based RNA-sequencing and bioinformatics strategy to analyze the lncRNAs of 9 samples under three different environment stresses (control, drought stress and re-watering, three replications), we totally identified 10,820 lncRNAs of high-confidence through five strict steps filtration, of which 9,989 were lincRNAs, 153 were inronic lncRNAs, 678 were anti-sense lncRNAs. Coding function analysis showed 6,470 lncRNAs may have the ability to code proteins. Small RNAs precursor analysis revealed that 196 lncRNAs may be the precursors to small RNAs, most of which (35.7%, 70) were miRNAs. Expression patterns analysis showed that most of lncRNAs were expressed at a low level and most inronic lncRNAs (75.95%) had a consistent expression pattern with their adjacent protein-coding genes. Further analysis of transcriptome data uncovered that lncRNAs XLOC_063105 and XLOC_115463 probably function in regulating two adjacent coding genes CotAD_37096 and CotAD_12502, respectively. Investigations of the content of plant hormones and proteomics analysis under drought stress also complemented the prediction. We analyzed the characteristics and the expression patterns of lncRNAs under drought stress and re-watering treatment, and found lncRNAs may be likely to involve in regulating plant hormones pathway in response to drought stress.
Conflict of interest statement
Figures

References
-
- Cheng J, Kapranov P, Drenkow J, Dike S, Brubaker S, Patel S, et al. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science. 2005; 308: 1149–1154. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

