Regulation of WNT Signaling by VSX2 During Optic Vesicle Patterning in Human Induced Pluripotent Stem Cells
- PMID: 27301076
- PMCID: PMC5156591
- DOI: 10.1002/stem.2414
Regulation of WNT Signaling by VSX2 During Optic Vesicle Patterning in Human Induced Pluripotent Stem Cells
Abstract
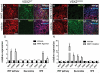
Few gene targets of Visual System Homeobox 2 (VSX2) have been identified despite its broad and critical role in the maintenance of neural retina (NR) fate during early retinogenesis. We performed VSX2 ChIP-seq and ChIP-PCR assays on early stage optic vesicle-like structures (OVs) derived from human iPS cells (hiPSCs), which highlighted WNT pathway genes as direct regulatory targets of VSX2. Examination of early NR patterning in hiPSC-OVs from a patient with a functional null mutation in VSX2 revealed mis-expression and upregulation of WNT pathway components and retinal pigmented epithelium (RPE) markers in comparison to control hiPSC-OVs. Furthermore, pharmacological inhibition of WNT signaling rescued the early mutant phenotype, whereas augmentation of WNT signaling in control hiPSC-OVs phenocopied the mutant. These findings reveal an important role for VSX2 as a regulator of WNT signaling and suggest that VSX2 may act to maintain NR identity at the expense of RPE in part by direct repression of WNT pathway constituents. Stem Cells 2016;34:2625-2634.
Keywords: Human induced pluripotent stem cells (hiPSCs); VSX2; WNT; optic vesicles.
© 2016 The Authors Stem Cells published by Wiley Periodicals, Inc. on behalf of AlphaMed Press.
Figures




Similar articles
-
Modeling human retinal development with patient-specific induced pluripotent stem cells reveals multiple roles for visual system homeobox 2.Stem Cells. 2014 Jun;32(6):1480-92. doi: 10.1002/stem.1667. Stem Cells. 2014. PMID: 24532057 Free PMC article.
-
The Role of FGF9 in the Production of Neural Retina and RPE in a Pluripotent Stem Cell Model of Early Human Retinal Development.Am J Ophthalmol. 2019 Oct;206:113-131. doi: 10.1016/j.ajo.2019.04.033. Epub 2019 May 10. Am J Ophthalmol. 2019. PMID: 31078532 Free PMC article.
-
RPE specification in the chick is mediated by surface ectoderm-derived BMP and Wnt signalling.Development. 2013 Dec;140(24):4959-69. doi: 10.1242/dev.096990. Epub 2013 Nov 13. Development. 2013. PMID: 24227655
-
Loss of MITF expression during human embryonic stem cell differentiation disrupts retinal pigment epithelium development and optic vesicle cell proliferation.Hum Mol Genet. 2014 Dec 1;23(23):6332-44. doi: 10.1093/hmg/ddu351. Epub 2014 Jul 9. Hum Mol Genet. 2014. PMID: 25008112 Free PMC article.
-
A regulatory loop involving PAX6, MITF, and WNT signaling controls retinal pigment epithelium development.PLoS Genet. 2012 Jul;8(7):e1002757. doi: 10.1371/journal.pgen.1002757. Epub 2012 Jul 5. PLoS Genet. 2012. PMID: 22792072 Free PMC article.
Cited by
-
BMP-induced reprogramming of the neural retina into retinal pigment epithelium requires Wnt signalling.Biol Open. 2017 Jul 15;6(7):979-992. doi: 10.1242/bio.018739. Biol Open. 2017. PMID: 28546339 Free PMC article.
-
Limitations and Promise of Retinal Tissue From Human Pluripotent Stem Cells for Developing Therapies of Blindness.Front Cell Neurosci. 2020 Sep 10;14:179. doi: 10.3389/fncel.2020.00179. eCollection 2020. Front Cell Neurosci. 2020. PMID: 33132839 Free PMC article. Review.
-
Wnt/β-Catenin Signaling Pathway Is Necessary for the Specification but Not the Maintenance of the Mouse Retinal Pigment Epithelium.Mol Cells. 2023 Jul 31;46(7):441-450. doi: 10.14348/molcells.2023.0029. Epub 2023 May 16. Mol Cells. 2023. PMID: 37190767 Free PMC article.
-
PAX6D instructs neural retinal specification from human embryonic stem cell-derived neuroectoderm.EMBO Rep. 2020 Sep 3;21(9):e50000. doi: 10.15252/embr.202050000. Epub 2020 Jul 23. EMBO Rep. 2020. PMID: 32700445 Free PMC article.
-
Chromatin Accessibility and Transcriptional Differences in Human Stem Cell-Derived Early-Stage Retinal Organoids.Cells. 2022 Oct 28;11(21):3412. doi: 10.3390/cells11213412. Cells. 2022. PMID: 36359808 Free PMC article.
References
-
- Chow RL, Lang RA. Early eye development in vertebrates. Annu Rev Cell Dev Biol. 2001;17:255–296. - PubMed
-
- Zuber ME, Gestri G, Viczian AS, et al. Specification of the vertebrate eye by a network of eye field transcription factors. Development. 2003;130:5155–5167. - PubMed
-
- Hyer J, Mima T, Mikawa T. FGF1 patterns the optic vesicle by directing the placement of the neural retina domain. Development. 1998;125:869–877. - PubMed
-
- Nguyen M, Arnheiter H. Signaling and transcriptional regulation in early mammalian eye development: a link between FGF and MITF. Development. 2000;127:3581–3591. - PubMed
-
- Vogel-Hopker A, Momose T, Rohrer H, et al. Multiple functions of fibroblast growth factor-8 (FGF-8) in chick eye development. Mech Dev. 2000;94:25–36. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

