Integrated Classification of Prostate Cancer Reveals a Novel Luminal Subtype with Poor Outcome
- PMID: 27302169
- PMCID: PMC5047668
- DOI: 10.1158/0008-5472.CAN-16-0902
Integrated Classification of Prostate Cancer Reveals a Novel Luminal Subtype with Poor Outcome
Abstract
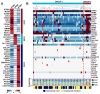
Prostate cancer is a biologically heterogeneous disease with variable molecular alterations underlying cancer initiation and progression. Despite recent advances in understanding prostate cancer heterogeneity, better methods for classification of prostate cancer are still needed to improve prognostic accuracy and therapeutic outcomes. In this study, we computationally assembled a large virtual cohort (n = 1,321) of human prostate cancer transcriptome profiles from 38 distinct cohorts and, using pathway activation signatures of known relevance to prostate cancer, developed a novel classification system consisting of three distinct subtypes (named PCS1-3). We validated this subtyping scheme in 10 independent patient cohorts and 19 laboratory models of prostate cancer, including cell lines and genetically engineered mouse models. Analysis of subtype-specific gene expression patterns in independent datasets derived from luminal and basal cell models provides evidence that PCS1 and PCS2 tumors reflect luminal subtypes, while PCS3 represents a basal subtype. We show that PCS1 tumors progress more rapidly to metastatic disease in comparison with PCS2 or PCS3, including PSC1 tumors of low Gleason grade. To apply this finding clinically, we developed a 37-gene panel that accurately assigns individual tumors to one of the three PCS subtypes. This panel was also applied to circulating tumor cells (CTC) and provided evidence that PCS1 CTCs may reflect enzalutamide resistance. In summary, PCS subtyping may improve accuracy in predicting the likelihood of clinical progression and permit treatment stratification at early and late disease stages. Cancer Res; 76(17); 4948-58. ©2016 AACR.
©2016 American Association for Cancer Research.
Conflict of interest statement
of Potential Conflicts of Interest: N. Erho is a bioinformatics group lead at GenomeDx Biosciences Inc. M. Alshalalfa is a bioinformatician at GenomeDx Biosciences Inc. H. Al-deen Ashab is a data scientist at Genomedx Biosciences Inc. E. Davicioni has ownership interest (including patents) in GenomeDx Biosciences Inc. R.J. Karnes reports receiving other commercial research support from GenomeDx Biosciences Inc. E.A. Klein has received speakers bureau honoraria from GenomeDx Biosciences Inc. A.E. Ross has ownership interest (including patents) in GenomeDx Biosciences Inc. Mandeep Takhar is a bioinformatician at GenomeDX. No potential conflicts of interest were disclosed by the other authors.
Figures




Comment in
-
Re: Integrated Classification of Prostate Cancer Reveals a Novel Luminal Subtype with Poor Outcome.J Urol. 2017 Mar;197(3 Pt 1):701-702. doi: 10.1016/j.juro.2016.12.032. Epub 2016 Dec 16. J Urol. 2017. PMID: 28208521 No abstract available.
-
A Systems Approach to Prostate Cancer Classification-Letter.Cancer Res. 2017 Dec 15;77(24):7131-7132. doi: 10.1158/0008-5472.CAN-16-3231. Epub 2017 Dec 6. Cancer Res. 2017. PMID: 29212852 No abstract available.
-
A Systems Approach to Prostate Cancer Classification-Response.Cancer Res. 2017 Dec 15;77(24):7133-7135. doi: 10.1158/0008-5472.CAN-17-0239. Epub 2017 Dec 6. Cancer Res. 2017. PMID: 29212854 No abstract available.
References
-
- Perner S, Demichelis F, Beroukhim R, Schmidt FH, Mosquera JM, Setlur S, et al. TMPRSS2:ERG fusion-associated deletions provide insight into the heterogeneity of prostate cancer. Cancer research. 2006;66(17):8337–41. - PubMed
-
- Tomlins SA, Laxman B, Dhanasekaran SM, Helgeson BE, Cao X, Morris DS, et al. Distinct classes of chromosomal rearrangements create oncogenic ETS gene fusions in prostate cancer. Nature. 2007;448(7153):595–9. - PubMed
-
- Singh D, Febbo PG, Ross K, Jackson DG, Manola J, Ladd C, et al. Gene expression correlates of clinical prostate cancer behavior. Cancer cell. 2002;1(2):203–9. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

