A Designed Experiments Approach to Optimizing MALDI-TOF MS Spectrum Processing Parameters Enhances Detection of Antibiotic Resistance in Campylobacter jejuni
- PMID: 27303397
- PMCID: PMC4885823
- DOI: 10.3389/fmicb.2016.00818
A Designed Experiments Approach to Optimizing MALDI-TOF MS Spectrum Processing Parameters Enhances Detection of Antibiotic Resistance in Campylobacter jejuni
Abstract
MALDI-TOF MS has been utilized as a reliable and rapid tool for microbial fingerprinting at the genus and species levels. Recently, there has been keen interest in using MALDI-TOF MS beyond the genus and species levels to rapidly identify antibiotic resistant strains of bacteria. The purpose of this study was to enhance strain level resolution for Campylobacter jejuni through the optimization of spectrum processing parameters using a series of designed experiments. A collection of 172 strains of C. jejuni were collected from Luxembourg, New Zealand, North America, and South Africa, consisting of four groups of antibiotic resistant isolates. The groups included: (1) 65 strains resistant to cefoperazone (2) 26 resistant to cefoperazone and beta-lactams (3) 5 strains resistant to cefoperazone, beta-lactams, and tetracycline, and (4) 76 strains resistant to cefoperazone, teicoplanin, amphotericin, B and cephalothin. Initially, a model set of 16 strains (three biological replicates and three technical replicates per isolate, yielding a total of 144 spectra) of C. jejuni was subjected to each designed experiment to enhance detection of antibiotic resistance. The most optimal parameters were applied to the larger collection of 172 isolates (two biological replicates and three technical replicates per isolate, yielding a total of 1,031 spectra). We observed an increase in antibiotic resistance detection whenever either a curve based similarity coefficient (Pearson or ranked Pearson) was applied rather than a peak based (Dice) and/or the optimized preprocessing parameters were applied. Increases in antimicrobial resistance detection were scored using the jackknife maximum similarity technique following cluster analysis. From the first four groups of antibiotic resistant isolates, the optimized preprocessing parameters increased detection respective to the aforementioned groups by: (1) 5% (2) 9% (3) 10%, and (4) 2%. An additional second categorization was created from the collection consisting of 31 strains resistant to beta-lactams and 141 strains sensitive to beta-lactams. Applying optimal preprocessing parameters, beta-lactam resistance detection was increased by 34%. These results suggest that spectrum processing parameters, which are rarely optimized or adjusted, affect the performance of MALDI-TOF MS-based detection of antibiotic resistance and can be fine-tuned to enhance screening performance.
Keywords: Campylobacter jejuni; MALDI-TOF MS; antibiotic resistance; antimicrobial resistance; designed experiments; spectrum processing.
Figures

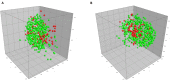
References
-
- Dieckmann R., Helmuth R., Erhard M., Malorny B. (2008). Rapid classification and identification of Salmonellae at the species and subspecies levels by whole-cell matrix-assisted laser desorption ionization-time of flight mass spectrometry. Appl. Environ. Microbiol. 74 7767–7778. 10.1128/AEM.01402-1408 - DOI - PMC - PubMed
-
- Fagerquist C. K., Garbus B. R., Williams K. E., Bates A. H., Boyle S., Harden L. A. (2009). Web-based software for rapid top-down proteomic identification of protein biomarkers, with implications for bacterial identification. Appl. Environ. Microbiol. 75 4341–4353. 10.1128/AEM.00079-79 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

