MYC/MIZ1-dependent gene repression inversely coordinates the circadian clock with cell cycle and proliferation
- PMID: 27339797
- PMCID: PMC4931031
- DOI: 10.1038/ncomms11807
MYC/MIZ1-dependent gene repression inversely coordinates the circadian clock with cell cycle and proliferation
Abstract
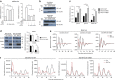
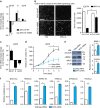
The circadian clock and the cell cycle are major cellular systems that organize global physiology in temporal fashion. It seems conceivable that the potentially conflicting programs are coordinated. We show here that overexpression of MYC in U2OS cells attenuates the clock and conversely promotes cell proliferation while downregulation of MYC strengthens the clock and reduces proliferation. Inhibition of the circadian clock is crucially dependent on the formation of repressive complexes of MYC with MIZ1 and subsequent downregulation of the core clock genes BMAL1 (ARNTL), CLOCK and NPAS2. We show furthermore that BMAL1 expression levels correlate inversely with MYC levels in 102 human lymphomas. Our data suggest that MYC acts as a master coordinator that inversely modulates the impact of cell cycle and circadian clock on gene expression.
Figures




References
-
- Dunlap J. C. Molecular bases for circadian clocks. Cell 96, 271–290 (1999). - PubMed
-
- Ukai H. & Ueda H. R. Systems biology of mammalian circadian clocks. Annu. Rev. Physiol. 72, 579–603 (2010). - PubMed
-
- Sahar S. & Sassone-Corsi P. Metabolism and cancer: the circadian clock connection. Nat. Rev. Cancer 9, 886–896 (2009). - PubMed
-
- Kelleher F. C., Rao A. & Maguire A. Circadian molecular clocks and cancer. Cancer Lett. 342, 9–18 (2014). - PubMed
-
- Filipski E. et al. Effects of chronic jet lag on tumor progression in mice. Cancer Res. 64, 7879–7885 (2004). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

