DDX3 Represses Stemness by Epigenetically Modulating Tumor-suppressive miRNAs in Hepatocellular Carcinoma
- PMID: 27344963
- PMCID: PMC4921922
- DOI: 10.1038/srep28637
DDX3 Represses Stemness by Epigenetically Modulating Tumor-suppressive miRNAs in Hepatocellular Carcinoma
Abstract
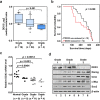
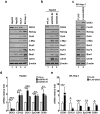
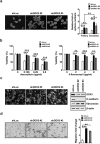
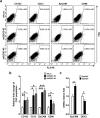
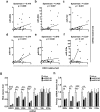
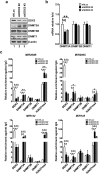
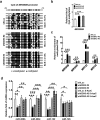
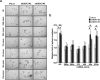
Studies indicate that the presence of cancer stem cells (CSCs) is responsible for poor prognosis of hepatocellular carcinoma (HCC) patients. In this study, the functional role of DDX3 in regulation of hepatic CSCs was investigated. Our results demonstrated that reduced DDX3 expression was not only inversely associated with tumor grade, but also predicted poor prognosis of HCC patients. Knockdown of DDX3 in HCC cell line HepG2 induced stemness gene signature followed by occurrence of self-renewal, chemoreisistance, EMT, migration as well as CSC expansion, and most importantly, DDX3 knockdown promotes tumorigenesis. Moreover, we found positive correlations between DDX3 level and expressions of tumor-suppressive miR-200b, miR-200c, miR-122 and miR-145, but not miR-10b and miR-519a, implying their involvement in DDX3 knockdown-induced CSC phenotypes. In addition, DDX3 reduction promoted up-regulation of DNA methyltransferase 3A (DNMT3A), while neither DNMT3B nor DNMT1 expression was affected. Enriched DNMT3A binding along with hypermethylation on promoters of these tumor-suppressive miRNAs reflected their transcriptional repressions in DDX3-knockdown cells. Furthermore, individual restoration of these tumor-suppressive miRNAs represses DDX3 knockdown-induced CSC phenotypes. In conclusion, our study suggested that DDX3 prevents generation of CSCs through epigenetically regulating a subset of tumor-suppressive miRNAs expressions, which strengthens tumor suppressor role of DDX3 in HCC.
Figures
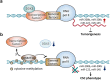
References
-
- Visvader J. E. & Lindeman G. J. Cancer stem cells in solid tumours: accumulating evidence and unresolved questions. Nat. Rev. Cancer 8, 755–768 (2008). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

