Upstream Open Reading Frames Differentially Regulate Gene-specific Translation in the Integrated Stress Response
- PMID: 27358398
- PMCID: PMC5016099
- DOI: 10.1074/jbc.R116.733899
Upstream Open Reading Frames Differentially Regulate Gene-specific Translation in the Integrated Stress Response
Abstract
Translation regulation largely occurs during initiation, which features ribosome assembly onto mRNAs and selection of the translation start site. Short, upstream ORFs (uORFs) located in the 5'-leader of the mRNA can be selected for translation. Multiple transcripts associated with stress amelioration are preferentially translated through uORF-mediated mechanisms during activation of the integrated stress response (ISR) in which phosphorylation of the α subunit of eIF2 results in a coincident global reduction in translation initiation. This review presents key features of uORFs that serve to optimize translational control that is essential for regulation of cell fate in response to environmental stresses.
Keywords: eukaryotic translation initiation; stress response; translation; translation control; translation regulation.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures



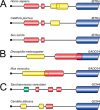
References
-
- Schwanhäusser B., Gossen M., Dittmar G., and Selbach M. (2009) Global analysis of cellular protein translation by pulsed SILAC. Proteomics 9, 205–209 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

