Development of Neutralization Assay Using an eGFP Chikungunya Virus
- PMID: 27367716
- PMCID: PMC4974516
- DOI: 10.3390/v8070181
Development of Neutralization Assay Using an eGFP Chikungunya Virus
Abstract
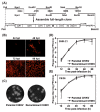
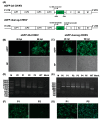
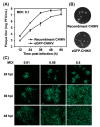
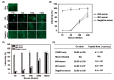
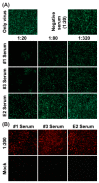
Chikungunya virus (CHIKV), a member of the Alphavirus genus, is an important human emerging/re-emerging pathogen. Currently, there are no effective antiviral drugs or vaccines against CHIKV infection. Herein, we construct an infectious clone of CHIKV and an eGFP reporter CHIKV (eGFP-CHIKV) with an isolated strain (assigned to Asian lineage) from CHIKV-infected patients. The eGFP-CHIKV reporter virus allows for direct visualization of viral replication through the levels of eGFP expression. Using a known CHIKV inhibitor, ribavirin, we confirmed that the eGFP-CHIKV reporter virus could be used to identify inhibitors against CHIKV. Importantly, we developed a novel and reliable eGFP-CHIKV reporter virus-based neutralization assay that could be used for rapid screening neutralizing antibodies against CHIKV.
Keywords: Chikungunya virus; antiviral; neutralization antibody; reporter virus.
Figures


Similar articles
-
Development of a neutralization assay based on the pseudotyped chikungunya virus of a Korean isolate.J Microbiol. 2020 Jan;58(1):46-53. doi: 10.1007/s12275-020-9384-0. Epub 2019 Nov 25. J Microbiol. 2020. PMID: 31768937 Free PMC article.
-
A potent neutralizing IgM mAb targeting the N218 epitope on E2 protein protects against Chikungunya virus pathogenesis.MAbs. 2015;7(6):1178-94. doi: 10.1080/19420862.2015.1083664. Epub 2015 Aug 25. MAbs. 2015. PMID: 26305993 Free PMC article.
-
Virus replicon particle based Chikungunya virus neutralization assay using Gaussia luciferase as readout.Virol J. 2013 Jul 15;10:235. doi: 10.1186/1743-422X-10-235. Virol J. 2013. PMID: 23855906 Free PMC article.
-
Development of Vaccines for Chikungunya Fever.J Infect Dis. 2016 Dec 15;214(suppl 5):S488-S496. doi: 10.1093/infdis/jiw271. J Infect Dis. 2016. PMID: 27920179 Free PMC article. Review.
-
Immune-Mediated Protection and Pathogenesis of Chikungunya Virus.J Immunol. 2016 Dec 1;197(11):4210-4218. doi: 10.4049/jimmunol.1601426. J Immunol. 2016. PMID: 27864552 Free PMC article. Review.
Cited by
-
Generation of a reporter yellow fever virus for high throughput antiviral assays.Antiviral Res. 2020 Nov;183:104939. doi: 10.1016/j.antiviral.2020.104939. Epub 2020 Sep 24. Antiviral Res. 2020. PMID: 32980446 Free PMC article.
-
Meeting report: the 4th symposium on animal models of non-human primates in Kunming, Yunnan, China.Zool Res. 2016 Nov 18;37(6):361-365. doi: 10.13918/j.issn.2095-8137.2016.6.361. Zool Res. 2016. PMID: 28105801 Free PMC article. No abstract available.
-
Generation and characterization of Japanese encephalitis virus expressing GFP reporter gene for high throughput drug screening.Antiviral Res. 2020 Oct;182:104884. doi: 10.1016/j.antiviral.2020.104884. Epub 2020 Aug 1. Antiviral Res. 2020. PMID: 32750466 Free PMC article.
-
Development of a novel Newcastle disease virus (NDV) neutralization test based on recombinant NDV expressing enhanced green fluorescent protein.Virol J. 2017 Nov 23;14(1):232. doi: 10.1186/s12985-017-0900-8. Virol J. 2017. PMID: 29169354 Free PMC article.
-
A single dose recombinant AAV based CHIKV vaccine elicits robust and durable protective antibody responses in mice.PLoS Negl Trop Dis. 2024 Nov 4;18(11):e0012604. doi: 10.1371/journal.pntd.0012604. eCollection 2024 Nov. PLoS Negl Trop Dis. 2024. PMID: 39495779 Free PMC article.
References
-
- Cruz D.J.M., Bonotto R.M., Gomes R.G.B., da Silva C.T., Taniguchi J.B., No J.H., Lombardot B., Schwartz O., Hansen M.A.E., Freitas-Junior L.H. Identification of novel compounds inhibiting Chikungunya virus-induced cell death by high throughput screening of a kinase inhibitor library. PLoS Neglect. Trop. Dis. 2013;7:181. doi: 10.1371/journal.pntd.0002471. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

