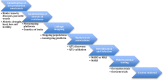
Genomic Tools in Cowpea Breeding Programs: Status and Perspectives
- PMID: 27375632
- PMCID: PMC4891349
- DOI: 10.3389/fpls.2016.00757
Genomic Tools in Cowpea Breeding Programs: Status and Perspectives
Abstract
Cowpea is one of the most important grain legumes in sub-Saharan Africa (SSA). It provides strong support to the livelihood of small-scale farmers through its contributions to their nutritional security, income generation and soil fertility enhancement. Worldwide about 6.5 million metric tons of cowpea are produced annually on about 14.5 million hectares. The low productivity of cowpea is attributable to numerous abiotic and biotic constraints. The abiotic stress factors comprise drought, low soil fertility, and heat while biotic constraints include insects, diseases, parasitic weeds, and nematodes. Cowpea farmers also have limited access to quality seeds of improved varieties for planting. Some progress has been made through conventional breeding at international and national research institutions in the last three decades. Cowpea improvement could also benefit from modern breeding methods based on molecular genetic tools. A number of advances in cowpea genetic linkage maps, and quantitative trait loci associated with some desirable traits such as resistance to Striga, Macrophomina, Fusarium wilt, bacterial blight, root-knot nematodes, aphids, and foliar thrips have been reported. An improved consensus genetic linkage map has been developed and used to identify QTLs of additional traits. In order to take advantage of these developments single nucleotide polymorphism (SNP) genotyping is being streamlined to establish an efficient workflow supported by genotyping support service (GSS)-client interactions. About 1100 SNPs mapped on the cowpea genome were converted by LGC Genomics to KASP assays. Several cowpea breeding programs have been exploiting these resources to implement molecular breeding, especially for MARS and MABC, to accelerate cowpea variety improvement. The combination of conventional breeding and molecular breeding strategies, with workflow managed through the CGIAR breeding management system (BMS), promises an increase in the number of improved varieties available to farmers, thereby boosting cowpea production and productivity in SSA.
Keywords: Vigna unguiculata; blackeye pea; cowpea; genomics; marker-assisted breeding.
Figures
Similar articles
-
A multi-parent advanced generation inter-cross (MAGIC) population for genetic analysis and improvement of cowpea (Vigna unguiculata L. Walp.).Plant J. 2018 Mar;93(6):1129-1142. doi: 10.1111/tpj.13827. Epub 2018 Feb 24. Plant J. 2018. PMID: 29356213
-
Introgression Breeding in Cowpea [Vigna unguiculata (L.) Walp.].Front Plant Sci. 2020 Sep 16;11:567425. doi: 10.3389/fpls.2020.567425. eCollection 2020. Front Plant Sci. 2020. PMID: 33072144 Free PMC article. Review.
-
Dissecting the Genetic Diversity of USDA Cowpea Germplasm Collection Using Kompetitive Allele Specific PCR-Single Nucleotide Polymorphism Markers.Genes (Basel). 2024 Mar 14;15(3):362. doi: 10.3390/genes15030362. Genes (Basel). 2024. PMID: 38540421 Free PMC article.
-
A major QTL corresponding to the Rk locus for resistance to root-knot nematodes in cowpea (Vigna unguiculata L. Walp.).Theor Appl Genet. 2016 Jan;129(1):87-95. doi: 10.1007/s00122-015-2611-0. Epub 2015 Oct 8. Theor Appl Genet. 2016. PMID: 26450274 Free PMC article.
-
Molecular genetics of race-specific resistance of cowpea to Striga gesnerioides (Willd.).Pest Manag Sci. 2009 May;65(5):520-7. doi: 10.1002/ps.1722. Pest Manag Sci. 2009. PMID: 19222045 Review.
Cited by
-
Development of Selection Indices for Improvement of Seed Yield and Lipid Composition in Bambara Groundnut (Vigna subterranea (L.) Verdc.).Foods. 2021 Dec 29;11(1):86. doi: 10.3390/foods11010086. Foods. 2021. PMID: 35010212 Free PMC article.
-
Characterization of Induced High Yielding Cowpea Mutant Lines Using Physiological, Biochemical and Molecular Markers.Sci Rep. 2020 Feb 28;10(1):3687. doi: 10.1038/s41598-020-60601-6. Sci Rep. 2020. PMID: 32111942 Free PMC article.
-
Quantitative Trait Loci Mapping for Yield and Related Traits in Cowpea.Genes (Basel). 2025 Feb 21;16(3):247. doi: 10.3390/genes16030247. Genes (Basel). 2025. PMID: 40149399 Free PMC article.
-
Cowpea Immature Pods and Grains Evaluation: An Opportunity for Different Food Sources.Plants (Basel). 2022 Aug 9;11(16):2079. doi: 10.3390/plants11162079. Plants (Basel). 2022. PMID: 36015383 Free PMC article.
-
Exploiting genetic and genomic resources to enhance productivity and abiotic stress adaptation of underutilized pulses.Front Genet. 2023 Jun 16;14:1193780. doi: 10.3389/fgene.2023.1193780. eCollection 2023. Front Genet. 2023. PMID: 37396035 Free PMC article. Review.
References
-
- Abate T., Alene A. D., Bergvinson D., Shiferaw B., Silim S., Orr A., et al. (2012). Tropical Grain Legumes in Africa and South Asia: Knowledge and Opportunities. Nairobi: International Crops Research Institute for the Semi-Arid Tropics.
-
- Agbicodo E. M., Fatokun C. A., Bandyopadhyay R., Wydra K., Diop N. N., Muchero W., et al. (2010). Identification of markers associated with bacterial blight resistance loci in cowpea [Vigna unguiculata (L.) Walp.]. Euphytica 175, 215–226. 10.1007/s10681-010-0164-5 - DOI
-
- Andargie M., Knudsen J. T., Pasquet R. S., Gowda B. S., Muluvi G. M., Timko M. P. (2014). Mapping of quantitative trait loci for floral scent compounds in cowpea (Vigna unguiculata L.). Plant Breed. 133, 92–100. 10.1111/pbr.12112 - DOI
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous